
INTERNATIONAL PROGRAMME ON CHEMICAL SAFETY
ENVIRONMENTAL HEALTH CRITERIA 47
SUMMARY REPORT ON THE EVALUATION OF SHORT-TERM TESTS
FOR CARCINOGENS (COLLABORATIVE STUDY ON IN VITRO TESTS)
This report contains the collective views of an international group of
experts and does not necessarily represent the decisions or the stated
policy of the United Nations Environment Programme, the International
Labour Organisation, or the World Health Organization.
Published under the joint sponsorship of
the United Nations Environment Programme,
the International Labour Organisation,
and the World Health Organization
World Health Orgnization
Geneva, 1985
The International Programme on Chemical Safety (IPCS) is a
joint venture of the United Nations Environment Programme, the
International Labour Organisation, and the World Health
Organization. The main objective of the IPCS is to carry out and
disseminate evaluations of the effects of chemicals on human health
and the quality of the environment. Supporting activities include
the development of epidemiological, experimental laboratory, and
risk-assessment methods that could produce internationally
comparable results, and the development of manpower in the field of
toxicology. Other activities carried out by the IPCS include the
development of know-how for coping with chemical accidents,
coordination of laboratory testing and epidemiological studies, and
promotion of research on the mechanisms of the biological action of
chemicals.
ISBN 92 4 154187 3
The World Health Organization welcomes requests for permission
to reproduce or translate its publications, in part or in full.
Applications and enquiries should be addressed to the Office of
Publications, World Health Organization, Geneva, Switzerland, which
will be glad to provide the latest information on any changes made
to the text, plans for new editions, and reprints and translations
already available.
(c) World Health Organization 1985
Publications of the World Health Organization enjoy copyright
protection in accordance with the provisions of Protocol 2 of the
Universal Copyright Convention. All rights reserved.
The designations employed and the presentation of the material
in this publication do not imply the expression of any opinion
whatsoever on the part of the Secretariat of the World Health
Organization concerning the legal status of any country, territory,
city or area or of its authorities, or concerning the delimitation
of its frontiers or boundaries.
The mention of specific companies or of certain manufacturers'
products does not imply that they are endorsed or recommended by the
World Health Organization in preference to others of a similar
nature that are not mentioned. Errors and omissions excepted, the
names of proprietary products are distinguished by initial capital
letters.
CONTENTS
SYNOPSIS - THE INTERNATIONAL PROGRAMME ON CHEMICAL SAFETY (IPCS):
COLLABORATIVE STUDY ON THE ASSESSMENT AND VALIDATION OF SHORT-TERM
TESTS FOR CARCINOGENS
1. SUMMARY REPORT ON IN VITRO TESTS
1.1. Introduction
2. THE COLLABORATIVE STUDY ON SHORT-TERM TESTS (CSSTT) 1981-83
3. CRITERIA FOR SELECTION OF THE TEST CHEMICALS
4. PURITY OF THE TEST CHEMICALS
5. CRITERIA FOR THE DEFINITION OF COMPLEMENTARY IN VITRO ASSAYS
FOR THE DETECTION OF POTENTIAL CARCINOGENS
6. ASSAYS AND END-POINTS
6.1. Bacteria
6.2. Fungi
6.3. Drosophila
6.4. Cultured mammalian cells
7. RESULTS
8. CONFIRMATION OF THE NON-MUTAGENICITY OF THE TEST CHEMICALS FOR
SALMONELLA
9. ASSESSMENT OF THE PERFORMANCE OF ASSAYS ON THE REDUCED LIST
9.1. Gene mutation in yeast
9.2. Drosophila somatic cell mutation assays
9.3. Assays for DNA damage SSB (single-strand breaks) and UDS
(detected via autoradiography or scintillation counting)
9.4. Assays for the induction of aneuploidy
9.5. Mammalian cell gene-mutation assays
9.6. Chromosomal-aberration assays
9.7. Assays for polyploidy induction
9.8. Sister chromatid exchange (SCE) assays
9.9. Transformation assays
10. SELECTION OF A PREFERRED COMPLEMENTARY ASSAY
11. CONCLUSIONS
REFERENCES
LIST OF PARTICIPANTS
Dr D. Amacher, Pfizer Central Research, Groton, Connecticut
Dr P. Arni, Ciba-Geigy, Basle, Switzerland
Dr J. Ashby, Central Toxicology Laboratory, Imperial Chemical
Industries, Ltd, Macclesfield, Cheshire, United Kingdom
Dr R. Baker, School of Public Health and Tropical Medicine,
University of Sydney, Sydney NSW, Australia
Dr J.C. Barrett, Laboratory of Pulmonary Function and Toxicology,
National Institute of Environmental Health Sciences, Research
Triangle Park, North Carolina, USA
Dr R.H. Barrett, The Boots Company Industrial Division,
Nottingham, United Kingdom
Dr M.O. Bradley, Merck, Sharp & Dohme, West Point, Pennsylvania,
USA
Dr T. Brooks, Shell Research, Ltd, Tunstall Laboratory, Kent,
United Kingdom
Dr A. Carere, Higher Institute of Health, Rome, Italy
Dr W. Caspary, National Toxicology Program, National Institute of
Environmental Health Sciences, Research Triangle Park, North
Carolina, USA
Dr D.V. Chitavichus, Institute of Medical Genetics, Moscow, USSR
Dr C.L. Crespi, Gentest Corporation, Woburn, Massachusetts
Dr N. Danford, Department of Genetics, University College of
Swansea, Swansea, United Kingdom
Dr B.J. Dean, Shell Research, Ltd, Tunstall Laboratory, Kent,
United Kingdom
Dr G. Delow, Paterson Laboratories, Christie Hospital & Holt Radium
Institute, Manchester, United Kingdom
Dr F.J. de Serres, National Institute of Environmental Health
Sciences, Research Triangle Park, North Carolina, USA
Dr G.R. Douglas, Environmental Health Centre, Department of
National Health and Welfare, Tunney's Pasture, Ottawa, Ontario,
Canada
Dr M. Draper, International Programme on Chemical Safety, World
Health Organization, Geneva, Switzerland
LIST OF PARTICIPANTS (contd.)
Dr E. Elmore, Northrop Services, Inc., Research Triangle Park,
North Carolina, USA
Dr L.R. Ferguson, Cancer Research Laboratory, Pathology
Department, The Medical School, Auckland, New Zealand
Dr K. Fujikawa, Drug Safety Evaluation Laboratories, Central
Research Division, Takeda Chemical Industries, Ltd., Osaka,
Japan
Dr R.C. Garner, Cancer Research Unit, University of York, York,
United Kingdom
Dr H. Glauert, Laboratory for Cancer Research, University of
Wisconsin Medical School, Madison, Wisconsin, USA
Dr D.K. Gulati, EHRT, Inc., Lexington, Kentucky, USA
Dr G. Hatch, Northrop Services, Inc., Research Triangle Park,
North Carolina, USA
Dr J. Heinisch, Institute for Microbiology, Darmstadt, Federal
Republic of Germany
Dr C. Howard, Central Toxicology Laboratory, Imperial Chemical
Industries, Ltd, Macclesfield, Cheshire, United Kingdom
Dr S. Inge-Vechtemov, Department of Genetics and Breeding,
Leningrad State University, Leningrad, USSR
Dr M. Ishidate, Division of Mutagenesis, National Institute of
Hygienic Sciences, Tokyo, Japan
Dr A. Knaap, Laboratory of Carcinogenesis and Mutagenesis,
National Institute of Public Health, Bilthoven, The
Netherlands
Dr T. Lakhanisky, Institut d'Hygiene et d'Epidemiologie,
Brussels, Belgium
Mr C.G. Lee, Chemical Defence Establishment, Porton Down,
Wiltshire, United Kingdom
Prof N. Loprieno, Institute of Biochemistry, Biophysics, and
Genetics, University of Pisa, Pisa, Italy
Dr D. McGregor, Development Toxicology, Inveresk Research Int.,
Ltd, Musselborough, Scotland
Dr B. Margolin, Biometry and Risk Assessment Program, National
Institute of Environmental Health Sciences, Research Triangle
Park, North Carolina
LIST OF PARTICIPANTS (contd.)
Dr C. Martin, Cancer Research Unit, University of York, York,
United Kingdom
Dr M. Mercier, International Programme on Chemical Safety, World
Health Organization, Geneva, Switzerland
Dr B.C. Myhr, Department of Genetics and Cell Biology, Litton
Bionetics, Inc., Kensington, Maryland
Dr A.J. Nelmes, Gallaher Limited, London, United Kingdom
Dr S. Nesnow, Carcinogenesis and Metabolism Branch, Health Effects
Research Laboratory, US Environmental Protection Agency,
Research Triangle Park, North Carolina
Dr E.R. Nestmann, Environmental Health Centre, Department of
National Health and Welfare, Tunney's Pasture, Ottawa, Ontario,
Canada
Dr G. Obe, Institute of General Genetics of the Free University of
Berlin, Berlin (West)
Dr T.J. Oberly, Lilly Research Laboratories, Greenfield Laboratory,
Greenfield, Indiana
Dr F. Palitti, Evolutionary Genetics Centre, Institute of Genetics,
Citta Universitaria, Rome, Italy
Dr S. Parodi, Scientific Institute for Tumours, University of
Genoa, Genoa, Italy
Dr J. Parry, Department of Genetics, University College of Swansea,
Singleton Park, Swansea, Wales, United Kingdom
Dr B.J. Phillips, British Industrial Biological Research
Association, Carshalton, Surrey, United Kingdom
Dr G. Probst, Lilly Research Laboratories, Greenfield Laboratory,
Greenfield, Indiana, USA
Mr C.R. Richardson, Central Toxicology Laboratory, Imperial
Chemical Industries, Ltd, Macclesfield, Cheshire, United Kingdom
Dr E. Matthews, Department of Molecular Biology, Litton Bionetics,
Inc., Kensington, Maryland
Dr T. Sanner, Laboratory for Environmental and Occupational Cancer,
Norsk Hydro's Institute for Cancer Research, Oslo, Norway
Dr M.D. Shelby, National Toxicology Program, National Institute of
Environmental Health Sciences, Research Triangle Park, North
Carolina
LIST OF PARTICIPANTS (contd.)
Dr J.W.I.M. Simons, State University of Leiden, Leiden, The
Netherlands
Dr J. Styles, Central Toxicology Laboratory, Imperial Chemical
Industries, Ltd, Macclesfield, Cheshire, United Kingdom
Dr W.A. Suk, Northrop Services, Inc., Research Triangle Park,
North Carolina
Dr G.F. van Went, Division of Toxicology and Chemical Analysis
of Foodstuffs, National Institute of Public Health, Bilthoven,
The Netherlands
Dr S. Venitt, Chemical Carcinogenesis Division, Pollards Wood
Research Station, Bucks, United Kingdom
Dr E. Vogel, Department of Radiation Genetics and Chemical
Mutagenesis, State University of Leiden, Leiden, The Netherlands
Dr R.C. von Borstel, Department of Genetics, The University of
Alberta, Alberta, Canada
Dr M.D. Waters, Genetic Toxicology Division, Health Effects
Research Laboratory, US Environmental Protection Agency,
Research Triangle Park, North Carolina, USA
Dr G. Williams, Naylor Dana Institute for Disease Prevention,
American Health Foundation, Valhalla, New York, USA
Dr F. Würgler, Institute of Toxicology, University of Zurich,
Schwerzenbach, Switzerland
Dr M.Z. Zdzienicka, Department of Radiation Genetics and Chemical
Mutagenesis, State University of Leiden, Leiden, The Netherlands
Dr E. Zeiger, Toxicology Research and Testing Program, National
Toxicology Program, National Institute of Environmental Health
Sciences, Research Triangle Park, North Carolina, USA
NOTE TO READERS OF THE CRITERIA DOCUMENTS
While every effort has been made to present information in the
criteria documents as accurately as possible without unduly
delaying their publication, mistakes might have occurred and are
likely to occur in the future. In the interest of all users of the
environmental health criteria documents, readers are kindly
requested to communicate any errors found to the Manager of the
International Programme on Chemical Safety, World Health
Organization, Geneva, Switzerland, in order that they may be
included in corrigenda, which will appear in subsequent volumes.
In addition, experts in any particular field dealt with in the
criteria documents are kindly requested to make available to the
WHO Secretariat any important published information that may have
inadvertently been omitted so that it may be considered in the
event of updating of the criteria document.
* * *
Partial financial support for the publication of this criteria
document was kindly provided by the United States Department of
Health and Human Services, through a contract from the National
Institute of Environmental Health Sciences, Research Triangle Park,
North Carolina, USA - a WHO Collaborating Centre for Environmental
Health Effects.
SYNOPSIS - THE INTERNATIONAL PROGRAMME ON CHEMICAL SAFETY (IPCS):
COLLABORATIVE STUDY ON THE ASSESSMENT AND VALIDATION OF SHORT-TERM
TESTS FOR CARCINOGENS
The first part of this project, dealing with in vitro studies,
has already been published by Elsevier, Amsterdam. The second
part, concerning in vivo studies, is expected to be completed and
evaluated by early 1985, with publication about one year later.
The rationale for the collaborative study was derived, to a
large extent, from the major findings of the "International Program
for the Evaluation of Short-Term Tests for Carcinogens" (IPESTTC)
(de Serres & Ashby, 1981). This study, in turn, arose from the
necessity to evaluate the efficacy of different short-term assays
proposed for supplementing the traditional long-term assay in the
rodent. The results of the IPESTTC clearly confirmed the value of
Salmonella reversion assays as suitable primary tests for potential
carcinogens and mutagens. However, it was also confirmed that some
known rodent carcinogens were either not detected, or only detected
with considerable difficulty, by such assays. The IPESTTC study
did not succeed in defining any complementary eukaryotic assay that
could be used to detect carcinogens, found to be negative in the
standard Salmonella reversion assay. Several assays showed
promise, but none could be recommended, because it was considered
that the supporting data base was too small.
It was against this background that the Collaborative Study on
the Assessment and Validation of Short-Term Tests for Genotoxicity
and Carcinogenicity (CSSTT) was proposed by the International
Programme on Chemical Safety (IPCS) and the National Institute of
Environmental Health Sciences (NIEHS) of the USA, as a
Participating Institution in the IPCS. The objective of the study
was to generate a wide range of test results, using a small group
of carefully-selected chemicals, which would contribute to an
empirical basis for the selection of one or more in vitro short-
term tests to complement the widely-used Salmonella test, developed
by Professor Bruce Ames.
Some 60 investigators presented nearly 90 individual sets of
assay results to the collaboration study, generating in all some
2500 dose-response relationships. Most of the currently available
in vitro eukaryotic assay systems were represented. The following
8 organic carcinogens known to be either inactive or difficult to
detect in the Salmonella assay were chosen: o-toluidine,
hexamethylphosphoramide (HMPA), safrole, acrylonitrile, benzene,
diethylhexylphthalate, phenobarbital, and diethylstilboestrol,
together with 2 chemicals, caprolactam and benzoin, for which there
was no evidence of carcinogenicity in 2-year, 2-species rodent
bioassays.
The results of the study were evaluated at a meeting of the
investigators, held at St. Simon's Island, Georgia, USA, on
22 - 28 October, 1983. Each group of assays was chaired by a
coordinator, and all the original data were discussed, evaluated,
and agreed. At these assay group meetings, important protocol
deficiencies were identified and these findings constitute one of
the most significant developments arising from the collaborative
study.
The findings indicate that carcinogens that are inactive or
difficult to detect in the Salmonella assay fall into 2 distinct
groups. The first group includes genotoxins that are probably non-
mutagenic to Salmonella because of deficiencies in the available
metabolic capacity of the assay system. Thus, HMPA, o-toluidine,
safrole, and acrylonitrile were detected by most of the eukaryotic
assays studied, indicating that there is a range of assays that can
complement the Salmonella mutation assay to a limited extent.
The other group of carcinogens, benzene, DEHP, DES, and
phenobarbital, displayed a more selective range of genotoxic
activities, and none of the assays was selectively sensitive to
them.
Of the assays studied, only the induction of chromosomal
aberrations, cell transformation, and gene mutation in mammalian
cells, and aneuploidy in yeast gave encouraging overall
performances for the 8 carcinogens, and, with the exception of the
first, formidable protocol deficiencies have to be remedied.
The 3 carcinogens, DES, phenobarbital, and DEHP, chosen to
represent the class of chemicals believed to induce tumours in
rodents without first modifying the integrity of the nuclear DNA,
each displayed a range of genotoxic activity. Thus, the term "non-
genotoxic" should be used only when a sufficiently large
genotoxicity data base has been established.
The collaborative study has provided considerable evidence to
support the view that in vitro assays should be classified as
confirmatory, complementary, and supplementary. This understanding
of the potential of an assay should provide a basis for the
elimination of redundancies in proposed combinations of test
procedures.
The major conclusion of the study was that the use of
chromosomal aberration assays, preferably in an agreed cell type,
in conjunction with an adequate assessment of the mutagenicity of a
chemical for Salmonella, might provide an efficient primary screen
for possible new carcinogens. A first priority should be the
application of resources to establish a generally acceptable and
applicable protocol for the conduct of this type of assay. The
adoption of a chromosomal-aberration assay as a common
complementary test has additional advantages in that it would allow
easy comparison of data, ready extension to supplementary
cytogenetic assays, and the provision of data derived from an
independent endpoint from the gene mutations of the Salmonella
assay. Finally, looking back at the developments in this field of
toxicology, it seems clear that to establish an assay at the level
of international acceptance requires about a decade of meticulous
scientific endeavour and international collaboration.
1. SUMMARY REPORT ON IN VITRO TESTS
1.1. Introduction
It was discovered in the late 1960s and early 1970s that many
chemicals underwent metabolic changes before they were capable of
inducing the processes leading to cancer. This led to the
development of in vitro techniques for bringing about such
metabolic transformations and, consequently, enormous progress was
made in the fields of mutagenesis and carcinogenesis. It soon
emerged that there was a strong correlation between the
carcinogenic activity of a chemical, particularly in rodents, and
its mutagenic properties, as demonstrated in a wide variety of in
vitro and in vivo experimental systems involving bacteria, yeasts,
insects, rodents, and mammalian cells in tissue culture. At the
same time, there was a growing awareness that some chemicals, for
example, vinyl chloride monomer, posed hitherto unsuspected health
dangers. The fact that increasing numbers of chemicals were being
shown to have toxic properties in the evolving in vitro
genotoxicity test systems, together with a series of disasters
associated with chemicals that resulted in considerable mortality
and morbidity, brought about a realization that appropriate
legislative control of chemicals was needed to ensure adequate
protection of human health. Thus, in the latter part of the 1970s,
there was an unprecedented amount of activity, both nationally and
internationally, in the field of chemical safety.
Public perception about the inadvertent exposure of human
beings to chemical carcinogens was greatly heightened by the
development of one particular assay system for the detection of
mutagens. This was the Salmonella typhimurium reversion test
incorporating a metabolic activation system, pioneered by Professor
Bruce Ames and his colleagues, and now known universally as the
Salmonella assay, or the Ames test. This assay system, which is
based on fundamental genetic and molecular biological principles,
produced test results within a week, and it was soon adopted by
scientists throughout the world, but more particularly in North
America and Europe. As a result, many hundreds of chemicals were
tested and pronounced on, as is now known, without full
appreciation of the technical difficulties of the test and the
biological significance of the results. The fact that many
chemicals in common use, ranging from food additives and cosmetics
to household products, were claimed to have mutagenic properties
and, hence, by implication could be carcinogens, received a
considerable amount of uncritical attention in the general
scientific and lay press.
It is generally accepted, and was so for the purposes of this
study, that the strongest evidence that a chemical is a carcinogen
is derived from either chemical or epidemiological findings in
human beings showing an unequivocal relationship between exposure
and the induction of malignant disease, supported by appropriate
animal studies; or, in the absence of adequate human exposure data,
the experimental induction in several rodent species of malignant
tumours following carefully-controlled systematic exposures to the
chemical for most of their life span. These rodent bioassays
require about 3 years of experimental effort to produce a
conclusion and are extremely costly. Thus, the prospect of
obtaining apparently equivalent information in a far shorter time
and at a fraction of the cost was immensely appealing. These hopes
were reinforced by the claims of confirmatory evidence provided by
an increasing number of different assay systems. It was these
developments that gave rise to the term "short-term tests" to refer
to assay systems that were believed to indicate carcinogenic
properties. It was even hoped that, in time, these assays would
replace, at least in part, the rodent bioassay. Although,
understandably, this idea received much uncritical support, many
scientists were sceptical about some of the claims made for these
assay systems, pointing out that there were many discrepancies in
the findings, so-called false positives and false negatives, and
there was much debate concerning the true predictive nature of
assay systems, singly and in combination, for rodent carcinogens
and the relevance of the findings for human disease.
By the early 1980s, many nations had adopted legislation to
control toxic chemicals, incorporating various test requirements
for acute and chronic effects and other preventive measures, such
as adequate labelling. However, the issue of a legal requirement
for specific short-term tests for mutagenic and carcinogenic
properties was usually avoided. In some cases, the problem was
recognized by the formulation of a discretionary set of
recommendations involving data from a battery or tier of assays in
which the Salmonella assay was a basic requirement. The enormous
international trade in chemicals and the realization that, for most
of the 50 000 or so chemicals in common use, little systematic
toxicological data existed, brought about an early appreciation of
the need to use scarce toxicological resources with maximum
efficiency and under international agreement. Because of the
profound implications for international trade in chemicals, the
Organization of Economic Cooperation and Development (OECD)
addressed the problem of ensuring that toxicological data developed
in one member state could be accepted in all member states. To
this end, an extensive series of guidelines for toxicity and other
testing was developed by international experts. Further, and of
great importance, a set of procedures was formulated to ensure good
laboratory practice and quality assurance. This major
international collaboration identified and codified the tests that
were believed to be necessary to provide sufficient data to ensure
safety in the use of a chemical. Obligatory measures and
procedures were set out to ensure that the tests were carried out
according to the highest standards, as laid down by international
agreement. The implementation of these guidelines and procedures
has proved to be a difficult and onerous task, even for well-
established toxicological laboratories. The attempt to set some of
the newly-developed short-term tests into a similar legislative
framework revealed many uncertainties, which it now appears can
only be resolved by international collaborative efforts on a scale
that has few precedents.
In the fields of genetics and molecular biology, from which the
science of mutagenesis has evolved, and which now have assumed
great importance for the understanding of carcinogenesis,
scientists have shown great interest and ingenuity in adapting the
particular biological systems they use for their research studies
to assay systems of possible general use. Unfortunately, what can
be a powerful and flexible tool in the hands of an experienced
research worker cannot easily be transformed into the somewhat
inflexible procedure that is required for a test system for routine
use throughout the world. Studies to evaluate the efficiency of
different short-term tests, which were started in a number of
countries more or less in parallel with the development of the
systems, were mostly concerned with the validity of the test
procedure. By the time legislative measures were being
consolidated in many countries, notably the Toxic Substances
Control Act, 1976, in the USA, legislators were faced with a
plethora of some 40 short-term tests, all claiming some promise for
revealing mutagenic or carcinogenic potential. Of considerable
concern to industry, at this time, was the desire of legislators to
have clear-cut criteria for yes/no decisions, even in a rapidly-
evolving subject such as carcinogenesis. If the Delaney clause
(United States Public Law, 1958), which states, in part, "that no
additive shall be deemed to be safe if it is found to induce cancer
when ingested by man or animal, or if it is found after tests which
are appropriate for evaluation of safety of food additives to
induce cancer in man or animals", were strictly applied to the
wider field of chemicals, the consequences for a society so
dependent on chemicals could be most serious.
It is understandable that, against this background, there was
considerable incentive and support for cooperative efforts to
resolve these important issues. The project, which is the subject
of this report, follows on from an earlier international study that
was based on the realization that, as a first step, the
effectiveness of any short-term test in discriminating between
carcinogens and noncarcinogens had to be established using, as
reference, chemicals for which extensive rodent bioassay results
were available.
The original project, called the International Programme for
the Evaluation of Short-Term Tests for Carcinogenicity (IPESTTC)
was carried out between 1977 and 1979 and 42 coded chemicals were
tested by over 60 scientists using some 30 assay systems. The
project arose as a result of initiatives by the Health and Safety
Executive and the Medical Research Council of the United Kingdom
and the National Institute of Environmental Health Sciences of the
USA. At the time of the planning of the IPESTTC, there was general
agreement concerning the value of the Ames test, which by this time
had a generally-accepted protocol. However, it had also become
apparent that certain carcinogens or classes of carcinogens failed
to mutate Salmonella; hence, there was a need to identify other
test systems that could complement the Ames assay. Thus, there
were 3 basic objectives in the programme. The first was to obtain
more systematic knowledge concerning the carcinogens that the Ames
test failed to detect. The second was to examine the ability of
selected test systems to discriminate between carcinogens and
noncarcinogens, and the third was to define assays that
complemented the bacterial mutation assays. To this end, 14
carcinogen/noncarcinogen pairs, together with 11 other carcinogens
and 3 other chemically-unrelated noncarcinogens, were specially
prepared with defined purity and distributed "blind" to the
investigators.
The principal results of the IPSSTTC, which have been published
in full (de Serres & Ashby, 1981), clearly confirmed the value of
the Salmonella assay as a suitable primary test for the detection
of potential mutagens and carcinogens. However, it was also
confirmed that some known rodent carcinogens were either not
detected, or only detected with difficulty, by this assay. The
study did not succeed in arriving at clear-cut conclusions
concerning a single complementary eukaryotic assay that was capable
of giving a positive response for the carcinogens found negative in
the standard Salmonella assay. Several assays that might serve in
this capacity were identified, but none was recommended for general
adoption, because it was considered that the supporting data base
was too small. An important practical aspect of the IPESTTC came
about through the meetings of investigators, where each assay group
discussed their results with immediate access to the raw data.
These discussions resolved discrepancies in the findings and
produced not only consensus views on the findings, but also
extremely valuable indications of protocol deficiencies, even for
the well-established Salmonella assay.
2. THE COLLABORATIVE STUDY ON SHORT-TERM TESTS (CSSTT) 1981-83
The IPESTTC was remarkably successful in attracting the
voluntary participation of a large number of scientists, together
with additional support from scientific institutions. The
conclusions from this study indicated clearly that priority should
be given to the identification of assay systems to complement the
Ames test. By 1981, the validity and usefulness of the Ames test
had been well substantiated, but it had also been established that
a number of important carcinogens were not detected, or detected
only with difficulty using this assay. It was, thus, generally
accepted that no single assay system could be relied on to detect
all carcinogens. This led to the proposal of the adoption of
testing schemes involving multiple in vitro assays in various
configurations such as batteries, tiers, or combinations of the
two. The basis for the selection and deployment of these multiple
tests was, and remains, theoretical prudence rather than empirical
evidence. That is, a variety of genetic end-points and organisms
representing different phylogenetic levels were selected with the
intent of not missing end-points or phylum-specific chemical
activity. Furthermore, in general, reliance was placed on the
deployment of genotoxicity assays, even though, by this time, other
factors were assuming importance in the biological etiology of
natural and chemically-induced cancer. Thus, the molecular targets
for investigation were no longer dominated by observations of
readily discernible changes in the sequence or integrity of nuclear
DNA, but involved consideration of subtle changes in chromosome,
gene, or oncogene function or expression (Klein, 1981; Reddy et
al., 1982; Tabin et al., 1982; Weiss, 1982). This implies that
some of the genetic end-points monitored in assay systems may
ultimately be shown not to be directly related to the critical
events in the etiology of some chemically-induced cancers (Cairns,
1981). Amidst these scientific controversies about the reliability
and biological significance of many short-term tests, registration
and health authorities were endeavouring to assess the genotoxic
data from the same short-term tests without adequate scientific
guidance for the interpretation of the all too frequently
discordant data.
It was against this background that the Collaborative Study on
Short-Term Tests for Genotoxicity and Carcinogenicity (CSSTT) was
proposed by the International Programme on Chemical Safety (IPCS)
and the National Institute of Environmental Health Sciences of the
USA, as a Participating Institution in the programme. The general
goals and designs of the study were outlined by an ad hoc Working
Groupa, which met at the invitation of the IPCS in Geneva, on
30 April - 1 May 1981. The plans were consolidated by an IPCS
Working Groupb, which met in Geneva, on 13 - 14 November 1981. The
--------------------------------------------------------------------
a Participants: Dr J. Ashby, Professor N.P. Bochkov, Dr B.E. Matter, Professor T.
Matsushima, Dr F.J. de Serres, Dr M. Shelby, and Professor F.H. Sobels.
b Participants: Dr J. Ashby, Dr G.R. Douglas, Dr M. Ishidate, Dr A. Leonard, Dr N.
Loprieno, Dr B.E. Matter, Professor T. Matsushima, Dr R. Montesano, Dr F.J. de Serres,
Dr M. Shelby, Professor F.H. Sobels, Dr M. Stoltz and Dr M. Waters.
subsequent coordination of the collaborative study was the
responsibility of a Steering Committee derived primarily from the
Working Group (J. Ashby; F. de Serres, Chairman; M. Ishidate Jr;
B. Margolin; B. Matter; M. Shelby; and M.H. Draper, IPCS).
The financial burden of the organization of this study was met
largely by the IPCS, together with some of its Participating
Institutions, particularly the National Institute of Environmental
Health Sciences in the USA. As in the IPESTTC project (de Serres &
Ashby, 1981), the funding of the assay work was provided, in the
majority of cases, by the individual investigators managing to
incorporate the work into their research programmes. This could
only occur with the goodwill and belief in the project of the
senior managements of the approximately 50 involved laboratories
from universities, research institutes, and industrial research
facilities, throughout the world. In addition, a number of
governments that support the IPCS provided financial assistance for
this study. These include the governments of Belgium, Italy, the
Netherlands, and, in particular, the United Kingdom.
The experience gained from the conduct of the IPESTTC and the
goodwill of the participants in that study were extensively drawn
on in the planning and organization of the CSSTT. The major
objective was defined as the generation of a wide range of test
results for a small group of carefully-selected rodent carcinogens
that would contribute to an empirical basis for selecting one or
more in vitro short-term tests as complementary to the Ames test.
The number of chemicals was kept to a minimum, because of the
experience of handling the 42 chemicals in the IPESSTC study. It
was argued that it was better to aim for an extensive data base for
a few chemicals than a reduced and patchy data base for a large
number. The chemicals were selected with much care, and those
chosen were all known to be particularly difficult to detect in
assay systems and, thus, would be expected to expose weaknesses and
inconsistencies in both the assay system and the protocols. This
indeed proved to be the case.
Some 60 investigators participating in the project carried out
nearly 90 individual sets of assays, generating, in all, some 2500
dose-response relationships. Most of the in vitro eukaryotic
tests, currently available, were represented. The 8 organic
carcinogens chosen as either inactive, or difficult to detect as
positive in the Salmonella assay, were: o-toluidine,
hexamethylphosphoramide, safrole, acrylonitrile, benzene,
diethylhexylphthalate, phenobarbital, and diethylstilboestrol,
together with 2 chemicals, caprolactam (Huff, 1982) and benzoin
(NTP, 1980), which had not shown any evidence of carcinogenicity in
2-year rodent bioassays. The criteria for the selection of these
chemicals is an important matter and the reasons for each inclusion
are given in the following section.
3. CRITERIA FOR THE SELECTION OF THE TEST CHEMICALS
Eleven carcinogens were defined as either difficult or
impossible to detect as bacterial mutagens in the IPESTTC study,
and 4 of these were selected for the CSSTT. These were
hexamethylphosphoramide, safrole, diethylstilboestrol, and
o-toluidine. The carcinogenicity of these agents for rodents is
well-established but only a small proportion of the Salmonella
assays conducted in the IPESTTC study detected them as mutagenic
(1/15, 4/17, 1/17, and 3/16, respectively). However, several of
these responses were not reproducible, and each was weak. This led
to the compounds being regarded as essentially non-mutagenic for
Salmonella in this study.
Hexamethylphosphoramide (HMPA)
Hexamethylphosphoramide (HMPA) is among the most potent of
animal carcinogens producing metastasizing nasal tumours in rats
exposed by inhalation.
There is a wealth of data indicating it to be non-mutagenic for
Salmonella, yet the results of the IPESTTC study suggested that it
was a general genotoxin in eukaryotic assays. One possible mode of
action for this agent is via the enzyme-mediated formation of
formaldehyde (Ashby & Lefevre, 1983). This is also a rat nasal
carcinogen that is difficult to detect as mutagenic in the
Salmonella assay but is a gene mutagen in human cells (Ashby &
Lefevre, 1983; Goldmacher & Thilly, 1983).
o-Toluidine
o-Toluidine is a relatively weak rodent hepatocarcinogen. Its
activity in this respect is interesting because it weakens the
earlier assumption that single ring aromatic amines, as opposed to
multiple ring arylamines such as 2-naphthylamine and 4-
aminobiphenyl, are non-carcinogenic. o-Toluidine was established
as difficult or impossible to detect in the Salmonella assay in
the IPESTTC study, which also showed that it was mutagenic to these
bacteria, if evaluated in the presence of norharman. These
collected findings suggested o-toluidine to be a general
genotoxin, which requires specific metabolic activation, rather
than an agent showing specificity of genetic action.
Safrole
Safrole is a weak rodent liver carcinogen and has been studied
extensively in the Salmonella assay. Although certain
investigators have reported it to be mutagenic, it is generally
found inactive in this assay. Both the alpha-acetoxy and the
sidechain epoxide derivatives are mutagenic, and these have been
suggested as the metabolites responsible for the carcinogenic
action observed. Safrole may, therefore, be a further example of a
general genotoxin that requires specific metabolic activation. Set
against this is the fact that it appears devoid of genetic activity
in vivo; thus, it gave a negative response in both the mouse bone-
marrow micronucleus assay and the in vivo rat liver unscheduled DNA
synthesis (UDS) assay. Consequently, the possibility cannot be
excluded that the tumours produced by this agent may be mediated
via some disturbance of normal homeostasis in the test animals
(i.e., by a non-genotoxic mechanism), despite its ability to induce
genetic changes in some in vitro test systems.
Diethylstilboestrol (DES)
Diethylstilboestrol (DES) is carcinogenic in both human beings
and experimental animals. It could have been selected for this
study simply on the basis of a recent paper that showed it to be
capable of transforming cells and inducing chromosomal damage in
the apparent absence of gene mutations (Barrett et al., 1981).
This finding was supported by the fact that, in the IPESTTC study,
DES was regarded by the investigators as a clastogen that was non-
mutagenic for Salmonella. Therefore, DES, together with benzene
(see below), were included in the study as agents that could
possibly demonstrate the reality of the genetic specificity of
action of some chemical carcinogens.
Benzene
Benzene is a unique carcinogen. Its possible leukaemogenic
activity in man has been discussed for many years, yet this effect
has been difficult to reproduce in animals. The compound is
nonetheless generally regarded as carcinogenic and extensive data
exist on its clastogenicity, particularly when evaluated in vivo.
Dean (1978) reviewed the literature on the genotoxicity of this
agent in short-term tests, and this, together with subsequent
studies, clearly defined it as non-mutagenic for bacteria. The
possibility of its complete inability to induce gene mutations in
vitro is implied in some papers, but its gene mutagenicity in vivo
has not yet been assessed.
Acrylonitrile
Similarities in structure between acrylonitrile and the
carcinogen vinyl chloride led Venitt to evaluate it for bacterial
mutagenicity. The debate that ensued in Mutation Research (Milvy &
Wolff, 1977; Venitt et al., 1977) regarding the mutagenic activity
of this agent in Salmonella and Escherichia coli can be summarized
by describing acrylonitrile as a chemical that could easily be
found non-mutagenic in a routine screening programme that employed
only bacteria as marker cells. The carcinogenicity of this agent
has been subsequently defined and reviewed. The question of
whether acrylonitrile interacts directly with DNA via a Michael
reaction, or via the intermediate metabolic formation of an epoxide
derivative, heightens interest in this agent.
Diethylhexylphthalate (DEHP)
Diethylhexylphthalate (DEHP) has been shown to produce
hepatomas in the rodent liver, yet the majority of experimental
data indicate it to be non-mutagenic for bacteria. It has been
proposed that the carcinogenicity of this agent is associated with
its ability to proliferate peroxisome microbodies in the rodent
liver (Moody & Reddy, 1978). This explanation would not require
DEHP itself to interact with nuclear DNA. The carcinogenicity of
DEHP has, therefore, been considered as possibly "epigenetic" in
origin, which increases the need to determine accurately its
genotoxic status in vitro. The extent to which DEHP is hydrolysed
to the corresponding mono-acid derivative (MEHP) could influence
the outcome of certain assays as the latter chemical, unlike the
former, is reported to be a clastogen and SCE-inducing agent in
vitro (Phillips et al., 1982; Tomita et al., 1982; Ashby, 1983).
Phenobarbital
Phenobarbital, although active as a rodent liver carcinogen,
also has significant tumour-promoting properties in the rodent
liver. In fact, the issue of whether phenobarbital is a pure
promoting agent devoid of cancer-initiating activity is of great
current interest. In contrast to DES, the rodent carcinogenicity
of phenobarbital appears not to be reflected in man, despite the
extensive and controlled exposure of epileptic patients (Clemmesen
& Hjalgrim-Jensen, 1977). Although this chemical is generally
regarded as non-genotoxic, limited evidence exists for its ability
to induce SCEs in vitro (Athanasiou & Kryrtopoulos, 1982; Ashby,
1983). This property may be related to its ionic composition (cf.
sodium saccharin, MEHP above, lacchaic acid, sodium benzoate, etc.,
for similar activity profiles) (Ashby, 1983). An additional point
of interest in this chemical is that Williams has presented data to
support the claim that phenobarbital is an example of an epigenetic
carcinogen (Williams, 1981).
The non-carcinogens caprolactam and benzoin
The selection of non-carcinogens suitable for use in the
evaluation of short-term tests has presented a stumbling block in
all validation exercises. In the early validation studies, non-
carcinogens were simply selected from compounds commonly regarded
as being non-carcinogenic. In some cases, no data existed
regarding their carcinogenicity, and this was taken as indicative
of inactivity. In the IPESTTC study, the non-carcinogens selected
were graded according to the extent and quality of the negative
data, and although this was an advance, the interpretation of
unexpected positive assay responses was difficult. This issue is
particularly important in relation to the widespread reference to
false-positive responses occurring in short-term tests; the
credibility that can be accorded to a false-positive response is
directly proportional to the certainty associated with the
compound's classification as a non-carcinogen. The fact that some
assumed non-carcinogens may eventually be classified as either weak
or organ-, strain-, sex-, or species-specific carcinogens might
lead to the re-evaluation of many previous examples of false-
positive assay responses.
In order to circumvent this problem in the current
collaborative study, particular attention was paid to the selection
of the 2 chemicals required to act as negative controls. The
agents selected were benzoin and caprolactam. The major criterion
for their selection was inactivity in recent cancer bioassays
conducted as part of the US National Toxicology Program. In the
reports of these studies (US NTP, 1980, 1982), it was concluded
that neither compound was carcinogenic in male or female Fischer
344 rats or B6C3Fl mice dosed at levels up to the maximum tolerated
dose over their lifetimes. These 2 studies were taken as
definitive as they represented the most detailed cancer bioassay
protocols currently in use. In addition, these agents were devoid
of overtly DNA-reactive substituents and were known to be non-
mutagenic for bacteria.
The 10 chemicals selected covered a wide range of structural
types and could, therefore, be considered representative of agents
encountered in the environment and chemical industries. In
addition, several of the carcinogens selected had been associated,
by other investigators, with possible mechanisms of cancer
induction other than the DNA-reaction/somatic mutation theory.
Finally, the 2 non-carcinogens were sufficiently well supported by
negative carcinogenicity data to ensure that clear decisions could
be made, regarding the significance of their genotoxic activity, in
vitro.
4. PURITY OF THE TEST CHEMICALS
As analytical techniques improve, it is possible to find trace
impurities in materials, formerly considered pure. Set against
this is the practical need to obtain large supplies of pure
chemicals for a study, such as the present collaborative study,
without inordinate costs and delays. This dilemma is heightened by
the history of the conduct of cancer bioassays where the test
chemical was often, if not usually, assumed to be pure in the
absence of appropriate analytical data. Many chemicals bioassayed
for carcinogenicity have been of technical quality and, therefore,
probably not more than 95% pure. Normally, this would not matter,
but when the cancer bioassay data are to be the ultimate reference
point, as in the present study, then the relative purity of the in
vitro test chemical becomes of importance. A 5% impurity, at high
doses, could be of biological significance. At one extreme, it can
be argued that material of similar purity (or impurity) to that
employed in the cancer bioassay should be assayed, but this may
lead to a further confounding of the total data base. At the other
extreme, it can be suggested that only absolutely pure materials
should be employed in vitro, whatever the cost and inconvenience
incurred in their preparation. This approach carries the penalty
that the carcinogenic response observed in mammals may have been
produced by impurities, in which case, activities observed in vitro
may not be correlated with carcinogenic activity (or inactivity).
This consideration is particularly relevant for benzene. The
most convincing carcinogenicity data for benzene were derived from
human beings exposed to it together with other chemicals, the
number and type of which varied from situation to situation. The
fact that the carcinogenicity of this chemical is difficult to
define in rodents has led to the suspicion that it may not be
benzene, but the chemicals used in association with it, that are
carcinogenic. Pure benzene was used in this study; a risk was
taken by doing so.
The purity criteria adopted for the present study entailed the
following assays of chemical purity:
a) One batch of each chemical of the highest grade commercial
samples available, usually 99% or more pure, was obtained.
b) The proton nuclear magnetic resonance spectrum, mass
spectrum, and infrared spectrum were determined and checked
for consistency with the proposed structure and for the
possible presence of impurities.
c) The elemental analysis (C, H, and N) was determined for
both liquids and solids; each was within 0.4% of the
theoretical value.
d) The melting point was determined and compared with
previously reported values for all solids. Because of
differences in thermometer calibrations, variations of less
than 4 °C were hard to interpret.
e) In 2 cases (safrole and o-toluidine), high-pressure liquid
chromatography (HPLC) was employed to evaluate trace
impurities seen by earlier assay methods.
f) Thin-layer chromatography (TLC) was undertaken on each
material, as appropriate. A variety of eluants and
detection systems was employed.
On the basis of the above determinations, the present chemicals
were deemed to be pure to a level of 99%. These techniques cannot
eliminate the chance that some activities observed for some of the
agents (both carcinogens and non-carcinogens) were due to
impurities. This admission is necessary, but is not exceptional,
given the paucity of analytical data usual in such studies,
including the reference cancer bioassays. Nonetheless, trace
impurities may have contributed to some activities, the weak gene
mutagenicity of phenobarbital in Salmonella being an example of
where further purification and reassaying in vitro might yield
useful additional data. Genotoxic impurities should not, however,
be too easily invoked to explain unexpected genotoxic responses.
First, similar concerns should apply to positive responses observed
in vitro for mammalian carcinogens, and second, such uncertainties
reflect equally on previous studies, the findings of which
constitute most of the established data base of this science. The
chemicals, made up in 5-g lots, were labelled and distributed to
the investigators in specially-sealed double containers. As these
chemicals were carcinogens and for various other reasons, they were
not distributed "blind".
5. CRITERIA FOR THE DEFINITION OF COMPLEMENTARY IN VITRO ASSAYS FOR
THE DETECTION OF POTENTIAL CARCINOGENS
In order to qualify as a complementary assay for routine use in
conjunction with the Salmonella plate-incorporation assay, a test
must have fulfilled the following requirements (Ashby et al.,
1983):
(a) It should have been successfully employed as a short-term
test in a number of laboratories, and should be
substantially represented in the literature.
(b) It should have performed well in the detection of the
present 8 carcinogens, while concomitantly finding both of
the non-carcinogens negative.
(c) Positive responses obtained with the 8 carcinogens tested
should have been unambiguous, dose-related, and
reproducible.
(d) Similar qualitative responses should have been observed by
the majority of the laboratories using the same assay.
(e) It should be appropriate for routine screening purposes,
i.e., not unduly demanding as far as resources and
technical facilities are concerned.
Four categories of assay may thus be defined:
(1) Assays suitable for general use in conjunction with the
Salmonella assay;
(2) Promising assays, i.e., assays that may be capable of
fulfilling criteria (a)-(e), but for which data are not
available for all of the test chemicals, or where repeat
studies are not available;
(3) Relatively new assays that, while not meeting criterion
(a), have performed well in the collaborative study, and
for which the present 10 chemicals form the greater part
of the available data base; these cases would best be
handled by the rapid and coordinated acquisition of
further information.
(4) Assays that are clearly inappropriate for routine use in
testing for potential carcinogens, i.e., that do not meet
criteria (b)-(e).
6. ASSAYS AND END-POINTS
As discussed above, the design of the present collaborative
study reflected the primary purpose of attempting to identify in
vitro eukaryotic assays, which are capable of detecting chemical
carcinogens, not readily detectable using bacterial assays. At the
organism level, 4 categories of assays were employed: bacteria,
yeast, fruit flies, and cultured mammalian cells. Within each of
these groups of organisms, a variety of test end-points were used.
Organisms and end-points will be described briefly and are
presented in Table 1. Full details are available in the published
assay working group reports and the reports of individual
investigators (Ashby et al., 1985).
Table 1. IPCS CSSTT test systems
-------------------------------------------------------------------
I. Bacteria
Salmonella typhimurium
TA97, TA98, TA100, TA102, HIS- HIS+
TA1535, TA1537, TA1538
TM677 AZAS AZAR
II. Fungi
Mutation:
Saccharomyces cerevisiae
XV185-14C ARG- ARG+; TRP- TRP+
HIS- HIS+; HOM- HOM+
RM52 HIS- HIS+
D7 ILV- ILV+
D6 and D61-M ADE- ADE+ ILV- ILV+
D5 small colonies due to mitochondrial
mutations
Schizosaccharomyces pombe
P1 red white colonies (ADE)
Aspergillus nidulans
35 methionine metabolism mutants
-------------------------------------------------------------------
Table 1. (contd.)
-------------------------------------------------------------------
II. Fungi (contd.)
Recombination:
Saccaromyces cerevisiae
JD1 gene conversion, tryptophan or
histidine prototrophy
D7 and D7-144 crossing-over, red and pink
colonies (ADE) gene conversion,
tryptophan prototrophy
PV-2 and PV-3 crossing-over, canavanine
resistance gene conversion, lysine
protrophy
D6 and D61-M crossing-over, cycloheximide
resistance
Aspergillus nidulans
P1 crossing over, green yellow
colonies
Aneuploidy:
Saccharomyces cerevisiae
D6 and D61-M red, cycloheximide sensitive white
cycloheximide resist
Aspergillus nidulans
P1 yellow sectors in green colonies
Illegitimate mating:
Saccharomyces cerevisiae
PV-4a and PV-4b mating type a
III. Drosophila
Somatic cell mutations
wing-mosaicism wing spots from mutations,
deletions, chromosome breakage,
mitotic recombination or aneuploidy
white-zeste eye eye spots from mutations or
mosaicism deletions
-------------------------------------------------------------------
Table 1. (contd.)
-------------------------------------------------------------------
III. Drosophila (contd.)
white/white coral eye spots (same events as wing
eye mosaicism spots above)
IV. Cultured mammalian cells
Metabolic cooperation
V79 survival of HGPRT- cells
Transformation
SHE colony assay
C3H10T1/2 focus assay
BALB/c 3T3 focus assay
SHE/SA7 viral enhancement of chemical
transformation-focus assay
RLV/FRE enhanced survival of Rauscher
leukaemia
virus-infected rat embryo cells
CHO invasive growth in agar
DNA damage
single-strand breaks
CHO alkaline sucrose sedimentation
rat hepatocytes alkaline elution
Unscheduled DNA synthesis
HeLa S3 scintillation counting-extraced
DNA
rat hepatocytes scintillation counting -DNA
extracted from isolated nuclei
rat hepatocytes autoradiography
Cytogenetic damage
chromosomal aberrations
CHO structural aberrations; micronuclei
Chinese hamster lung, structural aberrations; polyploidy
CHL
Chinese hamster, structural aberrations; polyploidy
liver,CHl-L aneuploidy
rat liver, RL4 structural aberrations; polyploidy
human lymphocytes structural aberrations
-------------------------------------------------------------------
Table 1. (contd.)
-------------------------------------------------------------------
Sister chromatid exchange
CHO
V79
rat liver, RL4
Gene mutations
L5178Y TK+/- TK-/-
OUAS OUAR
V79 HGPRT+ HGPRT-
V79 OUAS OUAR
CHO HGPRT+ HGPRT-
Human lymphoblasts
TK6 TK+/- TK-/-
AHH HGPRT+ HGPRT-
-------------------------------------------------------------------
6.1. Bacteria
The carcinogens included in the collaborative study were
selected on the basis of previously-published results indicating
their lack of activity in routinely-conducted Salmonella
mutagenicity tests. Five sets of Salmonella data were obtained in
this study to confirm the previous results and to provide bacterial
mutagenicity data on the batches of chemicals used in the current
study. Test data are reported for Salmonella typhimurium strains
TA97, TA98, TA100, TA102, and TA1535 in both pre-incubation and
plate incorporation protocols and TA1537 and TA1538 in the plate
assay only. S. typhimurium strain TM677 was used to detect
azaguanine resistant forward mutants, employing a treat and plate
method.
6.2. Fungi
Fungal systems, which offer the advantages of being both
microbial and eukaryotic, were used to evaluate a wide range of
genetic end-points. Test results from Saccharomyces cerevisiae,
Schizosaccharomyces pombe, and Aspergillus nidulans are reported in
relation to the following genetic end-points: nuclear gene
mutation (both forward and reverse), mitochondrial mutation, gene
conversion, mitotic crossing over, and aneuploidy.
6.3. Drosophila
Three separate laboratories reported test results from 3 newly-
developed assays for detecting genetic damage induced in somatic
cells of Drosophila. The white-zeste eye mosaicism test detects
eye spots resulting from mutations or deletions, while the wing
mosaicism and white/white coral eye mosaicism tests detect wing or
eye spots resulting from mutations, deletions, chromosome breakage,
mitotic recombination, or aneuploidy.
6.4. Cultured Mammalian Cells
Five major categories of chemically-induced effects were
reported for cultured mammalian cells: inhibition of metabolic
cooperation, transformation, DNA damage, cytogenetic effects, and
gene mutations.
Test data on the inhibition of metabolic cooperation, an assay
intended to detect promoting agents, as evidenced by increased
survival of HGPRT- V79 cells in the presence of an excess of HGPRT+
cells and 8-azaguanine or 6-thioguanine, were reported by 3
laboratories.
Six distinct transformation assays were reported including
those from 2 laboratories using Syrian hamster embryo (SHE) cells,
2 using C3H10T1/2 mouse cells, and single laboratories using the
Syrian hamster embryo/Simian adenovirus-7 and Rauscher leukaemia
virus-infected rat embryo cell assays. In addition, data for 5
compounds derived from a new assay in which the end-point was
invasive growth of CHO cells in soft agar were considered. Two
investigators, who had offered to generate data using the BHK21
transformation assay, withdrew from the study because of lack of
adequate time. This was disappointing as they had presented the
prospect of a link with the IPESTTC study in which the BHK21 assay
represented the sole transformation end-point.
The chemical induction of DNA single-strand breaks was
determined by assessing single-strand breaks using alkaline elution
or alkaline sucrose sedimentation. Tests for unscheduled DNA
synthesis were reported using protocols involving both
scintillation counting and autoradiography.
A large body of test data was reported for the 2 most commonly
used cytogenetic end-points, structural aberrations, and sister
chromatid exchanges. In addition, limited results were reported
for the induction of micronuclei, aneuploidy, and polyploidy.
Gene-mutation induction data were reported for 3 loci:
thymidine kinase (TK), hypoxanthine guanine phosphoribosyl
transferase (HGPRT), and NA+, K+ ATPase (Ouabain resistance) in
mouse, Chinese hamster, or human cells. These studies included 7
sets of test results from the L5178Y TK+/- system.
7. RESULTS
The investigators met at St. Simon's Island, Georgia, USA from
23 - 28 October 1983. During this meeting, each group of assay
participants, with the assay coordinator as chairman, discussed the
results with the raw data in front of them and individual results
were agreed. The group then formulated a consensus report on the
response of each chemical in the assay. These decisions were
incorporated into the coordinators' report, prepared during group
discussions on the overall performance of the assay and any defects
discovered. The coordinator's reports were presented and discussed
at plenary sessions, during which the conclusions and
recommendations of the study were developed. The coordinators
subsequently finalized their reports after further consultations
with the members of the group. The reports have been incorporated
into the text of the publication, which includes all the individual
reports of results as well as an editorial overview of the study
and a number of technical appendices (Ashby et al., 1985).
Table 2 includes, in summary form, all the agreed results for
each chemical in each test system in the study, as established in
the assay group discussions at St. Simon's Island. In order to
make some attempt at an overall assessment of assay performances,
the editorial group proceeded along the following lines. First,
the qualitative responses displayed in Table 2 were assumed to be
correct. Some of these results were unconfirmed and may, therefore,
represent false-positive or false-negative observations. Second,
it was decided that an assay should be capable of detecting at
least 2 of the selected carcinogenic test agents as positive,
before it could be assessed for possible use as a complementary
test. The extent to which inadequacies of individual test
protocols, as opposed to the insensitivity of the particular assay
or its genetic end-point, were responsible for negative responses
could only be discussed in cases where the same assay had been
conducted in 2 or more laboratories. Third, statistical
comparisons of the overall performance of assays were not
undertaken because, with the present rather unusual set of test
chemicals, this could yield meaningless if not misleading
conclusions, unless undertaken in depth. The entire data base was
entered into a computer file at NIEHS and detailed statistical
analyses may be undertaken, as appropriate.
In developing the discussion, it was further accepted that in
vitro assays are, by their constitution, only appropriate for the
identification of potential carcinogens; that is, in vitro tests
can be used to predict possible carcinogens, but not to define
them; at present, this can only be attempted by in vivo techniques.
It is, therefore, to be expected that certain agents will show
activity in vitro, but will be unable to express this potential in
vivo, because of their non-absorption, rapid excretion,
preferential detoxification, inappropriate partitioning, etc., in
mammals. In vitro assays cannot, and should not, be expected to
reveal these possibilities, which are, by definition, unique to
living animals. Activity seen in vitro for the present 2 non-
carcinogens was not used when assessing the overall performance of
the assays in question (Table 3), but rather, was used to emphasize
the true role and generic predictive weaknesses of in vitro assays.
On occasions, the assimilation of the large data base was made
easier by considering 4 carcinogens selected for the study as a
group (HMPA, o-toluidine, safrole, and acrylonitrile). This was
because it was known, at the outset, that they were more likely to
be detected by most assays, as each had already been established as
being genotoxic, though they were usually inactive in the
Salmonella mutation assay. The remaining carcinogens, with the
possible exception of benzene, were loosely regarded as non-
genotoxic, prior to this study. The collaborative study data base
generally supported the segregation of these 2 groups of
carcinogens and, thus, enabled a selective assessment of each assay
to be made. Some assays performed well with the first 4
carcinogens but poorly with the others, and some were insensitive
to this division and performed either generally well or poorly. A
possible further subdivision of the second 4 carcinogens became
evident as the review progressed and this will be discussed in the
section dealing in more detail with the assays.
Table 2. IPCS CSSTT in vitro study: summary of qualitative resultsa,b
--------------------------------------------------------------------------------
ASSAY ACN TOL HMPA SAF DES BEN PB DEHP ZOIN CAP
--------------------------------------------------------------------------------
1 BACTERIA
1.1.1 Salmonella ? N N N N N P N N N
1.1.2 Salmonella P N N N N N N N N N
1.1.3 Salmonella N N N N N N N N N N
1.1.4 Salmonella P P N N N N P N P N
1.1.5 Salmonella N N N N N N P N N N
2 FUNGI
2.1 Mutation
2.1.1 D7 N N P N N N N N N N
2.1.2 Asper 35 N N N N
2.1.3 D7 N N N N N P N N N N
2.1.4 XV185 P P P P P P P P P
2.1.5 XV185 N P P P P
2.1.6 P1 N N N P N N N N N N
2.1.7 D6 P N P N N N N N N N
2.1.8 D61-M P P P P N N N N N N
2.1.9 Mito. D5 P P N P N P N N N N
2.2 Gene conversion
2.2.1 D7 P N P N N N N P N N
2.2.2 D7 P N N P N N N N N N
2.2.3 D7-144 P P P P P P P P P
2.2.4 PV-3 N N N N N N N N N N
2.2.5 PV-2 N N N N N N N N N N
2.2.6 JD-1 P N N N N N N N N N
2.2.7 D7 P N P N N N N P N N
2.3 Crossing-over
2.3.1 D7 N N N N N N N N N N
2.3.2 Asper. 35 N N N N N N N N N N
2.3.3 D6 P N N N N P N N N
--------------------------------------------------------------------------------
Table 2 (contd.)
--------------------------------------------------------------------------------
ASSAY ACN TOL HMPA SAF DES BEN PB DEHP ZOIN CAP
--------------------------------------------------------------------------------
2.3.4 D61-M P N N N N N N N N N
2.3.5 D61-M P N N N N N N N N N
2.3.6 D7 N N N P N N N P P N
2.4 Aneuploidy
2.4.1 D6 P P P P P P P P N N
2.4.2 D61-M N N N N P P N N N ?
2.4.3 D61-M P P P P P P P P N N
2.4.4 Asper. 35 P N N P N N N N N N
3. DROSOPHILA SOMATIC CELLS
3.1.1 Wing spots P P P P N P N N N P
3.1.2 Eye spots P N P N N N N ? N P
3.1.3 Eye spots P P P P ? ? N ? N P
4. CULTURED MAMMALIAN CELLS (endpoints other than gene mutation)
4.1 Metabolic cooperation
4.1.1 V79 P P N ? N N ? P ? N
4.1.2 V79 P N N N N
4.1.3 V79 P N N ? N N N N N N
4.2 Transformation
4.2.1 BALB/C N N N N N N N N N N
4.2.2 C3H P P P P P ? ? P P P
4.2.3 C3H P ? p ? N
4.2.4 SHE P P P ? N P P P N ?
4.2.5 SHE P P P P P P N P N P
4.2.6 SHE/SA7 P ? N ? N ? N
4.2.7 Rl-FRE ? P N P P N N
4.2.8 CHO N ? N N N
4.3 Single-strand breaks
4.3.1 Rat Hepat. P P N P P N N N P N
4.3.2 CHO N P N P P P N N N
4.3.3 CHO P P N N P N N N N N
4.4 Unscheduled DNA synthesis (UDS)
4.4.1 Rat Hepat. (autorad) N N P N N N N N N N
4.4.2 Rat Hepat. (autorad) N N N N N N N N N N
4.4.3 Rat Hepat. (scint.) P P P P N P N P P N
4.4.4 HeLa (scint.) N P P P N N N N N N
4.4.5 HeLa (scint.) P ? P N N
4.5 Chromosomal aberrations
4.5.1 CHO P P N N P N P N N N
4.5.2 CHO N N P P N
4.5.3 CHO P N N P N
4.5.4 LYM P P P N P
4.5.5 CH1-L P P P N P N P N N N
4.5.6 CHL P P P P N P ? N P P
4.5.7 RL4 N P N N N N N N N N
4.6 Sister chromatid exchange
4.6.1 CHO P P P P N N N N N N
4.6.2 CHO P N N N N
4.6.3 CHO P N N P N
--------------------------------------------------------------------------------
Table 2 (contd.)
--------------------------------------------------------------------------------
ASSAY ACN TOL HMPA SAF DES BEN PB DEHP ZOIN CAP
--------------------------------------------------------------------------------
4.6.4 CHO N N N N N N
4.6.5 V79 P P P N N
4.6.6 RL4 N P P N N N N N N N
4.7 Micronucleus
4.7.1 CHO-MN P N N N N N N N N N
4.8 Polyploidy
4.8.1 CHL N P N N P N N N N P
4.8.2 CH1-L N N N N P N N N N N
4.8.3 RL4 N N N N P N N N N N
4.9 Aneuploidy
4.9.1 CH1-L aneupl. N P P N P P N P N N
4.9.2 CH1-L spindle N N N N P P P P N N
5. MAMMALIAN CELL MUTATIONS
5.1 L5178Y
5.1.1 L51-TK P P N P P P N N N N
5.1.2 L51-TK P N P N N P P P P N
5.1.3 L51-TK P N N N N N N N N N
5.1.4 L51-TK P N P P N N N N N N
5.1.5 L51-TK P P P ? P N P N P N
5.1.6 L51-TK ? P P P P ? N N
5.1.7 L51-TK N P ? N N
5.1.8 L51-OUA N N P P P P ? N ?
5.2 V79
5.2.1 V79-OUA N N N N N
5.2.2 L79-TG P P ? P P P ?
5.2.3 V79-TG N N N N N N N N P N
5.2.4 V79-TG N N N N N
5.3 CHO
5.3.1 CHO-TG N N N N N
5.3.2 CHO-OUA ? N N N N
5.4 Human lymphoblasts
5.4.1 Human lym. TK P P N P N N N N N N
5.4.2 Human lym. TG P P N N P P P N N N
--------------------------------------------------------------------------------
P = Positive
N = Negative
a Results of the study expressed by assay group and test agent. The
order of assays is as described in the Introduction and as displayed
in the associated key. The order of the chemicals is according to
decreasing genotoxicity, as shown in Fig. 1. The results shown are
those of individual investigators and may, therefore, differ from
those shown in the assay working group reports. Reproduced by
permission, from: Ashby et al. (1985).
b ACN - acrylonitrile. PB - phenobarbital.
TOL - o -toluidine. ZOIN - benzoin.
SAF - safrole. CAP - caprolactam.
BEN - benzene.
From the summarized total data base, as set out in Table 2, it
appears that the overall detection rates of the various assays were
poor, as evidenced by the predominance of negative results.
However, as one of the objectives was to challenge each assay
system with particularly difficult chemicals, in order to assess
its real status as a complementary test, this scatter of results
was to be expected. With only 8 "probes", it was not anticipated
that definitive answers would be obtained, but rather that systems
offering the best potential for development would be identified.
In the detailed examinations of the raw data carried out by the
investigators at St. Simon's Island, many of the discrepancies in
the findings were associated with differences in protocol. Matters
such as dose levels, sampling times, statistical methods, cell
lines, and metabolic competence of the systems all proved to be
critical aspects of protocol. Had there been agreement on all
these aspects, it is highly probable that the results would have
been far more consistent. Thus, as with the IPESTTC, the CSSTT had
again demonstrated the importance of protocol detail in assay
systems as the key to interlaboratory consistency. Such protocol
development is a definite and obviously difficult process in the
evolution of a test system, and the CSSTT has clearly demonstrated
that, with the exception of the Ames test, this has not been
achieved satisfactorily for any of the possible complementary assay
systems. However, at least a possible mechanism for bringing about
significant improvements is emerging.
Table 3. Overall performance of assays for 8 carcinogensa,b
---------------------------------------------------------------------------------------------------------------------
Class of assay/ Summary of qualitative results expressed as positive tests/total tests Overall
chemical ACN TOL HMPA SAF Overall DES BEN PB DEHP Overall ZOIN CAP Overall performance
for the 8
carcinogens
---------------------------------------------------------------------------------------------------------------------
2. Fungi
2.1 Mutation 4/4 3/5 4/5 4/5 15/19 0/3 2/4 1/4 1/4 4/15 2/5 2/5 4/10 50%
(79%) (27%) (40%)
2.4/4.9.1 3/5 3/5 3/5 3/5 12/20 4/5 4/5 2/5 3/5 13/20 0/5 0/5 0/10 62%
Aneuploidy (60%) (65%) (0%)
3. Drosophila 3/3 2/3 3/3 2/3 10/12 0/3 1/3 0/3 0/3 1/12 0/3 3/3 3/6 46%
somatic cells (83%) (8%) (50%)
4. Cultured mammalian cells
4.3 Single- 2/3 3/3 0/3 2/3 7/12 3/3 1/3 0/2 0/3 4/11 1/3 0/3 1/6 48%
strand breaks (58%) (36%) (17%)
4.4 UDS 1/2 3/3 2/3 3/3 9/11 0/2 1/3 0/2 1/2 2/9 1/2 0/3 1/5 55%
(scintillation) (82%) (22%) (20%)
4.5 Chromosomal 4/4 3/5 3/5 2/4 12/18 3/4 3/6 3/4 0/3 9/17 1/4 2/6 3/10 60%
aberrations (67%) (53%) (30%)
4.6 SCE 2/3 4/5 3/4 2/4 11/16 0/2 0/5 1/3 0/2 1/12 0/2 0/5 0/7 43%
(69%) (8%) (0%)
5. Mammalian cell 6/7 5/10 7/10 5/10 23/37 5/8 6/10 4/9 1/8 16/35 3/9 0/8 3/17 55%
mutations (L51, (62%) (46%) (18%)
V79 human cells)
---------------------------------------------------------------------------------------------------------------------
Table 3. (contd.)
---------------------------------------------------------------------------------------------------------------------
Class of assay/ Summary of qualitative results expressed as positive tests/total tests Overall
chemical ACN TOL HMPA SAF Overall DES BEN PB DEHP Overall ZOIN CAP Overall performance
for the 8
carcinogens
---------------------------------------------------------------------------------------------------------------------
Overall 25/31 26/39 25/38 23/37 15/30 18/39 11/32 6/30 8/33 7/38
activity (81%) (67%) (66%) (62%) (50%) (46%) (34%) (20%) (24%) (18%)
---------------------------------------------------------------------------------------------------------------------
99/145 33/69 17/62 15/72
(68%) (48%) (27%) (21%)
----------------------- ----------------------- -----------
132/214 32/133
(62%) (24%)
------------------------------------------------------------------------------
149/276
(54%)
--------------------------------------------------------
---------------------------------------------------------------------------------------------------------------------
1. Salmonella 2/5 1/5 0/5 0/5 3/20 0/5 0/5 3/5 0/5 3/20 1/5 0/5 1/10 15%
(15%) (15%) (10%)
4.2 Transformation 3/3 5/6 4/6 3/6 15/21 3/5 2/5 1/3 4/5 10/18 1/4 276 3/10 64%
(71%) (55%) (30%)
---------------------------------------------------------------------------------------------------------------------
a The responses shown in the reduced data base (Table 4) are represented as number of positive responses/number of
observations made. Questionable responses have been eliminated from the numerator but included in the denominator.
The carcinogen sensitivity of each class of assay has been calculated, but not their sensitivity (as only 2
carcinogens were employed), nor their accuracy (because of the unique handling of non-carcinogens in this study).
The responses of the Salmonella assay and the transformation assays are shown at the foot of the Table.
b ACN - acrylonitrile TOL - o-toluidine SAF - safrole
BEN - benzene PB - phenobarbital ZOIN - benzoin
CAP - caprolatam
8. CONFIRMATION OF THE NON-MUTAGENICITY OF THE TEST CHEMICALS FOR
SALMONELLA
The most fundamental assumption made at the outset of the
collaborative study was that the 8 carcinogens selected were either
difficult or impossible to detect as positive using the standard
Salmonella mutation assay, and that the 2 non-carcinogens would be
equally inactive. This assumption was based, in part, on the
results of the IPESTTC and partly on a general perception of the
published literature, available earlier in 1981, on chemicals
picked in November, 1981.
As the collaborative study progressed, it was decided to
re-evaluate the test chemicals in the Salmonella assay as an
integral part of the study. This was triggered by a variety of
factors. First, the detailed literature review undertaken by The
Environmental Mutagen Information Centre (EMIC) revealed reports on
the mutagenicity of some of the chemicals for Salmonella. Second,
the use of the pre-incubation test was becoming increasingly
common, and not all of the agents had been tested using this
protocol. Third, 2 new strains of Salmonella were announced by
Professor Bruce Ames at that time (TA97 and TA102) (Ames et al.,
1975), and the possible activity of these chemicals became of
interest. Finally, these 10 agents had not been tested in parallel
before, nor had any common criteria been applied for the assessment
of their relative mutagenicity or chemical purity.
The bacterial study included both the plate-incorporation and
pre-incubation assay protocols, a range of S9 mixes and the 7 major
strains of Salmonella, including TA97 and TA102. The Salmonella
forward mutation system of Skopek et al. (de Serres & Ashby, 1981)
was included for purposes of comparison (strain TM 677; assay
1.1.2), and Zeiger (assay 1.1.4) employed uninduced hamster as well
as induced rat S9 in his experiments because this was his standard
practice (Table 2).
Negative conclusions were recorded in 347 of 360 tests, the
exceptions being listed below:
o-toluidine weak activity in TA1535 and TA100, in 1 out of
5 laboratories, and only when using hamster S9.
acrylonitrile weak activity in TA1535 and TA100 (+S9), in one
laboratory, weak activity in TM677, in another
laboratory (-S9), and questionable activity in
TA102, in a third laboratory (-S9).
benzoin weak activity in TA1535 (-S9) and questionable
activity in TA100 (-S9), in 1 laboratory out of
5.
phenobarbital S9 independent weak activity in TA1535, in 2
laboratories, and in TA100 in 1 laboratory.
Questionable activity was also seen in TA100
(S9) in one of the laboratories recording
activity in TA1535.
HMPA, benzene, safrole, caprolactam, DEHP, and DES showed no
evidence of mutagenic activity.
The assay working group concluded that these data confirm that
the present 10 test chemicals are either difficult or impossible to
detect as bacterial mutagens using the routinely-employed test
protocols of the Salmonella assay; thus, their selection for the
present study was endorsed.
9. ASSESSMENT OF THE PERFORMANCE OF THE ASSAYS ON THE REDUCED LIST
Application of the criteria set out in section 7 for the
overall assessment of an assay performance as complementary to the
Ames test eliminates about half the data in Table 2. The remaining
data sets are presented in Table 4. As the major conclusions of
the CSSTT are based on this "reduced" data base, some justification
for the elimination principles employed is necessary. The
Salmonella data, by definition of complementary assays, have not
been included. The decision to remove certain classes of assay
from consideration, because they were not yet suitably developed
for routine use, or had proved difficult to establish in
independent laboratories, was a decision of the editors; however,
their views were usually supported by the conclusions of the
appropriate assay group reports. Thus, the transformation assays
were eliminated, largely on the basis of the conclusions of the
working group. However, it is relevant that, in laboratories where
certain of these assays were performing reliably, they appeared to
provide an efficient complementary assay. The metabolic
cooperation assays were similarly eliminated, as these did not
appear to be optimal for use as a complementary test. The removal
of certain classes of assay, which were generally insensitive to
the present carcinogens, was automatically justified by the aims of
this study. The rat hepatocyte autoradiographic UDS assays and the
CHO micronucleus test were eliminated on the basis of this
principle.
Table 4. IPCS CSSTT in vitro study: summary of reduced data basea,b
--------------------------------------------------------------------------------
ASSAY ACN TOL HMPA SAF DES BEN PB DEHP ZOIN CAP
--------------------------------------------------------------------------------
2 FUNGI
2.1 Mutation
2.1.4 XV185 P P P P P P P P P
2.1.5 XV185 N P P P P
2.1.7 D6 P N P N N N N N N N
2.1.8 D61-M P P P P N N N N N N
2.1.9 Mito. D5 P P N P N P N N N N
2.4 Aneuploidy
2.4.1 D6 P P P P P P P P N N
2.4.2 D61-M N N N N P P N N N ?
2.4.3 D61-M P P P P P P P P N N
2.4.4 Asper. 35 P N N P N N N N N N
3. DROSOPHILA SOMATIC CELLS
3.1.1 Wing spots P P P P N P N N N P
3.1.2 Eye spots P N P N N N N ? N P
3.1.3 Eye spots P P P P ? ? N ? N P
4. CULTURED MAMMALIAN CELLS (endpoints other than gene mutation)
4.3 Single-strand breaks
4.3.1 Rat Hepat. P P N P P N N N P N
4.3.2 CHO N P N P P P N N N
4.3.3 CHO P P N N P N N N N N
--------------------------------------------------------------------------------
Table 4 (contd.)
--------------------------------------------------------------------------------
ASSAY ACN TOL HMPA SAF DES BEN PB DEHP ZOIN CAP
--------------------------------------------------------------------------------
4.4 Unscheduled DNA synthesis (UDS)
4.4.3 Rat Hepat. (scint.) P P P P N P N P P N
4.4.4 HeLa (scint.) N P P P N N N N N N
4.4.5 HeLa (scint.) P ? P N N
4.5 Chromosomal aberrations
4.5.1 CHO P P N N P N P N N N
4.5.2 CHO N N P P N
4.5.3 CHO P N N P N
4.5.4 LYM P P P N P
4.5.5 CH1-L P P P N P N P N N N
4.5.6 CHL P P P P N P ? N P P
4.5.7 RL4 N P N N N N N N N N
4.6 Sister chromatid exchange
4.6.1 CHO P P P P N N N N N N
4.6.2 CHO P N N N N
4.6.3 CHO P N N P N
4.6.4 CHO N N N N N N
4.6.5 V79 P P P N N
4.6.6 RL4 N P P N N N N N N N
4.9 Aneuploidy
4.9.1 CH1-L aneupt. N P P N P P N P N N
5. MAMMALIAN CELL MUTATIONS
5.1 L5178Y
5.1.1 L51-TK P P N P P P N N N N
5.1.2 L51-TK P N P N N P P P P N
5.1.4 L51-TK P N P P N N N N N N
5.1.5 L51-TK P P P ? P N P N P N
5.1.6 L51-TK ? P P P P ? N N
5.1.7 L51-TK N P ? N N
5.1.8 L51-OUA N N P P P P ? N ?
5.2 V79
5.2.2 L79-TG P P ? P P P ?
5.4 Human lymphoblasts
5.4.1 Human lym. TK P P N P N N N N N N
5.4.2 Human lym. TG P P N N P P P N N N
--------------------------------------------------------------------------------
P = Positive
N = Negative
a Results from selected assays shown in Table 2. This reduction in the
data base is justified in the text and forms the basis for the
selection of a generally-applicable complementary in vitro assay.
Reproduced by permission, from: Ashby et al. (1985).
b ACN - acrylonitrile. PB - phenobarbital.
TOL - o-toluidine. ZOIN - benzoin.
SAF - safrole. CAP - caprolactam.
BEN - benzene.
Elimination of individual assays that failed to detect any or
only 1 of the test chemicals, in cases where all 10 had been
evaluated, requires separate justification. In most of these
cases, other investigators using nominally the same assay detected
several of the carcinogens; the inference is, therefore, that the
particular protocol used was more at fault than the class of assay.
The individual assays eliminated by this criterion are listed
below, and it is clear that different underlying reasons may have
led to their poor performance.
Six assays only detected acrylonitrile as positive. The fact
that this carcinogen was the most generally genotoxic of the 8
carcinogens tested suggests that the sensitivity of the assay
protocols in question was too low, rather than that the assay class
as a whole was of no potential value as a complementary assay.
However, 4 of these assays formed part of 2 classes eliminated for
reasons of general insensitivity (yeast gene conversion and yeast
crossing over). These 6 data sets were:
2.2.3
yeast conversion
2.2.6
2.3.4
yeast crossing over
2.3.5
4.7.1 CHO micronucleus
5.1.3 L5178Y TK gene mutation
The ability of 2 of the polyploidy assays (4.8.2 and 4.8.3) to
detect only DES may reflect the exceptional potency of this agent
as a spindle poison. DES may, therefore, be too potent an agent to
act solely as the monitor for the sensitivity of aneuploidy assays.
Five assays gave isolated positive responses that appeared to
have no explanation. These effects may, therefore, reflect the
technical false-positive responses of these tests. Whatever the
reason, the weakness of these responses, together with the
generally negative context in which they occurred, suggests that
these activities should only be related with caution to the
carcinogenicity of the test agents. These activities involved the
following chemicals and assays:
benzoin assay 5.2.3 (V79 TG gene mutation)
o-toluidine assay 4.5.7 (RL4 chromosomal aberrations)
HMPA assay 4.4.1 (hepatocyte autorad. UDS)
assay 2.2.1 (yeast D7 gene mutation)
benzene assay 2.1.3 (yeast D7 gene mutation)
The above decisions to eliminate certain assays and data were
designed to optimize the relevance of the conclusions from the
present collaborative study. The steps outlined removed from
consideration assays that were not optimally performed or that were
representative of a class of assay generally deemed unsuitable for
routine adoption, as a complementary assay at this stage of their
development. Similar exclusion principles have been adopted by the
several GENETOX review groups, and the resultant data base provides
a more reliable reflection of the current stature of individual
assays. The positive effect that the above decisions had on the
CSSTT data base is demonstrated in Fig. 1. From this, it is
evident that the composite sensitivity of the assays to the 8
carcinogens has been increased without any corresponding loss of
specificity (see boxed area of Fig. 1). This figure also
demonstrates that the degree of genotoxicity of the present 10
chemicals, relative to each other, was not affected by the
elimination procedures employed, and thus provides a compelling
justification for proceeding with a detailed assessment of the data
shown in Table 4.
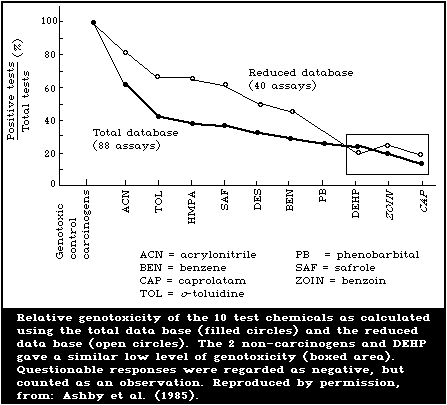 The performance of assays on the reduced list was carefully
assessed by the Editorial Committee, as it was felt that, for the
guidance of the reader, some overall evaluation of this complex
project was necessary, even though at this stage the opinions were
those of scientists, closely associated with the project, and,
thus, might be subjective. Close attention was paid to the
conclusions drawn by the assay groups regarding the usefulness of
individual assays. However, not all of these working groups
approached an assessment of their assays within the context
outlined originally. On occasion, the recommendations and
conclusions derived below are at variance with conclusions of the
group but, generally, they are consistent with them or represent
the only ones available.
9.1. Gene Mutation in Yeast
These assays are, phylogenically, the nearest to the Salmonella
assay and share a similar genetic end-point. Previous studies have
also established that genotoxic agents, found active in Salmonella,
are generally also active in the corresponding yeast gene-mutation
assays. It was, therefore, expected that these 2 classes of assay
would share similar sensitivities. Fourteen positive responses
were observed within the group comprising HMPA, o-toluidine,
safrole, and acrylonitrile, while only 3 positive responses were
observed for benzene, DEHP, DES, and phenobarbital, and there was
general agreement on the gene mutagenicity of HMPA (4/5 positive)
(Table 3, Fig. 2). The enhanced sensitivity of the yeast gene-
mutation assays, as opposed to the Salmonella mutation assay, to
the 4 assumed genotoxins may be due to endogenous metabolism,
especially evident when the cells are in active growth. It was,
therefore, concluded that this class of assay might represent a
possible alternative to the Salmonella assay, and might present
advantages, on occasion, because of its enhanced sensitivity to
certain genotoxins. The mitochondrial mutation assay (2.1.9) has
been included in this discussion, but should correctly be regarded
as a distinct assay. Two potential problems are presented by these
assays, and by yeast assays in general. First, they are not yet
conducted according to an agreed protocol. For example, the data
shown in Table 2 represent the consensus gained after conducting
assays within certain laboratories under a range of test
conditions: low or high glucose, stationary, semi-stationary, and
log-phase cells, etc. In addition, the largely positive data base
of assay 2.1.4 includes 2 classes of positive response (see assay
group report and investigator report). Second, the Salmonella
assay must remain the preferred primary assay for detecting gene
mutagens in vitro, owing to the extensive data base available and
the large number of laboratories involved (ca 2000).
It was concluded that appropriate yeast assays present a useful
method of detecting gene mutagens, and, on occasion, can be more
sensitive than the Salmonella test. However, given the current
state of the art, they would not be recommended as primary gene-
mutation assays and their usefulness as a single complement to the
Salmonella test is reduced by the repetition of the genetic end-
point involved. The preferential detection by these assays of
HMPA, o-toluidine, safrole, and acrylonitrile is referred to later.
9.2. Drosophila Somatic Cell Mutation Assays
New somatic cell mutation assays have recently been developed
in Drosophila. Their novelty is evidenced by the fact that data
supporting their general sensitivity to genotoxic agents (i.e.,
agents readily detected by the Salmonella assay) are only now
being collected for publication. General sensitivity is, however,
claimed (see working group report). Two different types of assay
were represented. Assays 3.1.1 and 3.1.3, although based in
different tissues, were capable of detecting a range of induced
genetic events, i.e., chromosomal breakage, deletions and
aneuploidy, gene mutations, and mitotic recombinations. These 2
assays were, in fact, more sensitive to the present test chemicals
than assay 3.1.2, where an inappropriate route of administration
was employed and the range of detected mutagenic events was limited
to gene mutations and deletions. Nevertheless, the general
concordance between the responses recorded in each of these 3
assays suggests that they are likely to yield reproducible data
between laboratories.
The performance of assays on the reduced list was carefully
assessed by the Editorial Committee, as it was felt that, for the
guidance of the reader, some overall evaluation of this complex
project was necessary, even though at this stage the opinions were
those of scientists, closely associated with the project, and,
thus, might be subjective. Close attention was paid to the
conclusions drawn by the assay groups regarding the usefulness of
individual assays. However, not all of these working groups
approached an assessment of their assays within the context
outlined originally. On occasion, the recommendations and
conclusions derived below are at variance with conclusions of the
group but, generally, they are consistent with them or represent
the only ones available.
9.1. Gene Mutation in Yeast
These assays are, phylogenically, the nearest to the Salmonella
assay and share a similar genetic end-point. Previous studies have
also established that genotoxic agents, found active in Salmonella,
are generally also active in the corresponding yeast gene-mutation
assays. It was, therefore, expected that these 2 classes of assay
would share similar sensitivities. Fourteen positive responses
were observed within the group comprising HMPA, o-toluidine,
safrole, and acrylonitrile, while only 3 positive responses were
observed for benzene, DEHP, DES, and phenobarbital, and there was
general agreement on the gene mutagenicity of HMPA (4/5 positive)
(Table 3, Fig. 2). The enhanced sensitivity of the yeast gene-
mutation assays, as opposed to the Salmonella mutation assay, to
the 4 assumed genotoxins may be due to endogenous metabolism,
especially evident when the cells are in active growth. It was,
therefore, concluded that this class of assay might represent a
possible alternative to the Salmonella assay, and might present
advantages, on occasion, because of its enhanced sensitivity to
certain genotoxins. The mitochondrial mutation assay (2.1.9) has
been included in this discussion, but should correctly be regarded
as a distinct assay. Two potential problems are presented by these
assays, and by yeast assays in general. First, they are not yet
conducted according to an agreed protocol. For example, the data
shown in Table 2 represent the consensus gained after conducting
assays within certain laboratories under a range of test
conditions: low or high glucose, stationary, semi-stationary, and
log-phase cells, etc. In addition, the largely positive data base
of assay 2.1.4 includes 2 classes of positive response (see assay
group report and investigator report). Second, the Salmonella
assay must remain the preferred primary assay for detecting gene
mutagens in vitro, owing to the extensive data base available and
the large number of laboratories involved (ca 2000).
It was concluded that appropriate yeast assays present a useful
method of detecting gene mutagens, and, on occasion, can be more
sensitive than the Salmonella test. However, given the current
state of the art, they would not be recommended as primary gene-
mutation assays and their usefulness as a single complement to the
Salmonella test is reduced by the repetition of the genetic end-
point involved. The preferential detection by these assays of
HMPA, o-toluidine, safrole, and acrylonitrile is referred to later.
9.2. Drosophila Somatic Cell Mutation Assays
New somatic cell mutation assays have recently been developed
in Drosophila. Their novelty is evidenced by the fact that data
supporting their general sensitivity to genotoxic agents (i.e.,
agents readily detected by the Salmonella assay) are only now
being collected for publication. General sensitivity is, however,
claimed (see working group report). Two different types of assay
were represented. Assays 3.1.1 and 3.1.3, although based in
different tissues, were capable of detecting a range of induced
genetic events, i.e., chromosomal breakage, deletions and
aneuploidy, gene mutations, and mitotic recombinations. These 2
assays were, in fact, more sensitive to the present test chemicals
than assay 3.1.2, where an inappropriate route of administration
was employed and the range of detected mutagenic events was limited
to gene mutations and deletions. Nevertheless, the general
concordance between the responses recorded in each of these 3
assays suggests that they are likely to yield reproducible data
between laboratories.
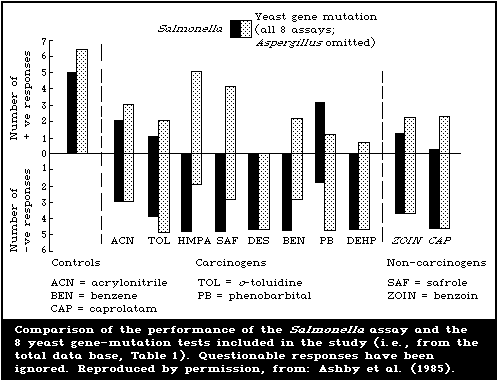 Assays using Drosophila require minimal technical facilities
but may prove relatively time-consuming. They present the appeal
of conducting an assay on a multicellular organism with its
inherent metabolic capability. It cannot be automatically assumed
that this capability is significantly related to that of a mammal,
but the sensitivity of these assays to a range of genotoxic agents
is consistent with their possession of a broad range of metabolic
pathways.
The 2 most sensitive assays (3.1.1 and 3.1.3) detected 5 and
perhaps 6 of the test agents, one of which was the non-carcinogen
caprolactam (see subsequent discussion of CAP and ZOIN). Activity
was mainly centred in the first group of carcinogens (Table 4, Fig.
3). This performance suggests that somatic mutation in Drosophila,
especially the assays capable of detecting a wide range of genetic
events, presents a promising new assay. However, their
preferential detection of HMPA, o-toluidine, safrole, and
acrylonitrile reduces their value as a general complementary assay
(cf. similar discussion of yeast gene-mutation assays, SCE assays,
and final conclusions). The present discussion is only related to
the use of these assays for predicting the possible mammalian
carcinogenicity of an agent; their use for predicting in vivo
mammalian mutagenic events should be considered separately (see
also discussions of mammalian cell gene-mutation assays).
Assays using Drosophila require minimal technical facilities
but may prove relatively time-consuming. They present the appeal
of conducting an assay on a multicellular organism with its
inherent metabolic capability. It cannot be automatically assumed
that this capability is significantly related to that of a mammal,
but the sensitivity of these assays to a range of genotoxic agents
is consistent with their possession of a broad range of metabolic
pathways.
The 2 most sensitive assays (3.1.1 and 3.1.3) detected 5 and
perhaps 6 of the test agents, one of which was the non-carcinogen
caprolactam (see subsequent discussion of CAP and ZOIN). Activity
was mainly centred in the first group of carcinogens (Table 4, Fig.
3). This performance suggests that somatic mutation in Drosophila,
especially the assays capable of detecting a wide range of genetic
events, presents a promising new assay. However, their
preferential detection of HMPA, o-toluidine, safrole, and
acrylonitrile reduces their value as a general complementary assay
(cf. similar discussion of yeast gene-mutation assays, SCE assays,
and final conclusions). The present discussion is only related to
the use of these assays for predicting the possible mammalian
carcinogenicity of an agent; their use for predicting in vivo
mammalian mutagenic events should be considered separately (see
also discussions of mammalian cell gene-mutation assays).
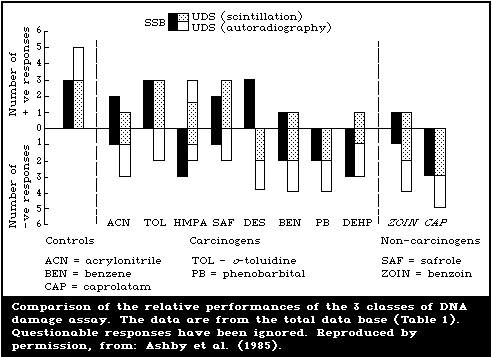 9.3. Assays for DNA Damage SSB (Single-Strand Breaks) and UDS
(Detected via Autoradiography or Scintillation Counting)
Although both of these classes of assay are designed to be
sensitive to the consequences of the primary interaction of the
test chemical (or a metabolite) with DNA, there are good reasons
for not considering them as equivalent. These reasons are
discussed in the assay working group report and are supported by
aspects of the present data base (Tables 2 and 3). For example,
HMPA was inactive in the 3 SSB assays yet was active in the assays
for UDS (2+ and 1?). In contrast, DES was uniformly active in the
3 SSB assays and inactive in the UDS tests. The fact that
unanimity between these 2 assay groups can be achieved is evidenced
by the responses seen for TOL (6/6+), SAF (5/6+), and PB (4/4-).
The relative sensitivity of these 3 classes of assay are shown in
Fig. 3. It should be noted that some legislative authorities have
discussed the need for assay data indicating primary interaction of
a test chemical with DNA. Their requirement could be met with data
generated from either of these groups of assay, i.e., the 2 are
generally regarded as being different methods of assaying a similar
phenomenon. As can be seen, this may not be so.
Perhaps one of the most surprising findings of the CSSTT is the
insensitivity demonstrated by the 2 autoradiographic assays (4.4.1
and 4.4.2, eliminated from Table 4). These were conducted
according to similar protocols and are well represented in earlier
literature. Their poor performance in the present study suggests
that they are unsuitable for use as a complementary assay, though
their use for confirming the activity in mammalian cells of
previously defined bacterial mutagens remains. Another intriguing
aspect of their poor performance is that it provides evidence to
suggest that the autoradiographic end-point is less sensitive than
the other end-points under review (with the exception of the
activity seen for HMPA in one assay, 4.4.1). This insensitivity is
unlikely to be due to metabolic incompetence as the hepatocytes
employed were primary isolates, the metabolic competence of which
had been previously established. Furthermore, the possibility that
this divergence of data was due to these agents producing only
short-patch repair is reduced, given the range of chemically
different carcinogens employed. Whether some of the positive
responses observed in the SSB assays were a direct result of cell
necrosis is discussed below.
Assays for SSB and UDS would be expected to be primarily
sensitive to the 4 confirmed genotoxins represented here, HMPA,
o-toluidine, safrole, and acrylonitrile, and this was observed (cf.
Table 3 and the discussion of the yeast and Drosophila gene-
mutation assays in this section). The only confirmed success with
the remaining 4 carcinogens was the ability of all of the SSB
assays to detect DES. Non-reproducible activity was seen in both
classes of assay for benzene (2/6), benzoin (2/5), and DEHP (1/5).
The latter isolated positive response was strongly endorsed by the
investigator (4.4.3) and requires further investigation.
A topic for future research is the extent to which cell
necrosis, with the consequent formation of broken DNA, contributes
to the outcome of SSB assays. For example, the isolated positive
response observed for benzene (assay 4.3.2) was only observed at
higher dose levels than those employed by the other 2 investigators
in this group. This response may, therefore, represent a technical
false-positive response induced solely by DNA strand breakage as
the result of toxicity. Equally, the extent to which
"mitotic" DNA synthesis has been successfully blocked during the
conduct of a scintillation UDS assay relates directly to the
credibility accorded to weak induced effects. The above 2 concerns
may explain some of the divergencies of data evident within these 3
classes of assay for the present chemicals.
Given the simple requirement to provide evidence of the ability
of a chemical to induce primary DNA damage in mammalian cells, the
investigator is faced with the following decisions:
(a) If the agent is already defined as a bacterial mutagen,
any of the 3 classes of assay (SSB or UDS via
autoradiography or scintillation counting) discussed
herein could be used.
(b) If the agent is inactive as a bacterial mutagen, use of
either the SSB or scintillation counted UDS assays could
present advantages, depending on the chemicals involved.
The results obtained for HMPA and DES suggest that the use
of both assays should be considered, if an assay for "DNA
damage" is required as a complement to bacterial
mutagenicity.
It was, therefore, concluded that the simple legislative
requirement for "evidence of DNA damage" was too vague to be
meaningful. If complementary data are being sought, the
autoradiographic end-point should be avoided and data should be
derived from an SSB assay and/or scintillation counted UDS assay.
The failure of the CSSTT to indicate clearly a single class of
assay, within this group, for routine adoption suggests that
attention should be given to developing the sensitivity of DNA
damage assays to the point that only a single test is required.
The relative performance of these 3 classes of DNA repair assays is
shown in Fig. 3.
9.4. Assays for the Induction of Aneuploidy
In the past, assays for the induction of aneuploidy have not
generally been considered as predictive tests for carcinogenicity.
In the first place, few appropriate in vitro assays existed, and
secondly, the induction of aneuploidy was assumed to have greater
relevance for the promotional, as opposed to the initiatory stages
of carcinogenesis. Thus, while the previous data base has included
the evaluation of a few reference genotoxins such as MNU, it has
mainly focused on cancer-promoting agents such as the phorbol
esters and saccharin, etc. Two investigators have provided the
greater part of the present data base for this class of assay by
presenting data derived from yeast assays (2.4.1.3). In addition,
data derived using mammalian cells were presented (4.9.1) together
with a set of data observed in Aspergillus (2.4.4).
The performance of the aneuploidy assays in the CSSTT was
impressive (Table 2 and 3). In the case of the 2 sets of yeast
data provided from one laboratory (2.4.1 and 2.4.3), a 100%
correlation between the reported carcinogenicity of the test agents
and their activity in vitro was observed. In particular, the
unanimous detection of both DEHP and phenobarbital by these
aneuploidy assays is unique in this study.
The primary conclusion regarding this group of assays is,
therefore, that they present high promise for development as a
class of complementary assays. The development phase should be
influenced by the following considerations, some of which are of
sufficient importance to ensure that they are resolved before any
particular aneuploidy assay is recommended for general adoption.
(a) The findings of the CSSTT suggest that the induction of
aneuploidy may be a critical step in the mechanism of
action of complete carcinogens. These observations
require independent confirmation in other laboratories,
for both the yeast and the mammalian cell systems.
(b) In the case of the D61-M yeast data from the 2
laboratories considered, major discrepancies were evident.
These were all of the same nature and represented the
failure of one laboratory to detect anything beyond
benzene and DES. It is possible that this divergence was
due to technical factors affecting the mitotic or
metabolic activity of the yeast. For example, evidence
from the fungal studies supported an earlier observation
that growing cells are more sensitive to genotoxins than
stationary cells. This could be because of the reduced
metabolic activity of stationary cells. Thus, this
divergence might be due to a relatively minor technical
consideration, but it must be defined, understood,
corrected, and defended in the literature before any
general adoption of yeast aneuploidy assays can be
recommended. The question of whether the D6 strain can be
regarded as repetitive of the D61-M strain also requires
resolution; if it is to be retained, its efficacy must be
confirmed in other laboratories.
(c) The isolated set of Aspergillus data suggests that this
assay is relatively insensitive. Clearly, if the
induction of aneuploidy is to be pursued as a useful
complementary end-point, and if Aspergillus nidulans is to
be selected as the marker organism, a separate data base
will have to be generated to justify this choice.
(d) Requirements similar to those in (c) must apply to the
isolated set of aneuploidy data generated in the Chinese
hamster liver fibroblast assay (4.9.1). This assay is
relatively novel; thus, apart from an expansion of the
data base and the commissioning of independent studies,
the following point requires attention. The cells used in
this study were of between the 9 - 15th passage. The
corollary for using such a spread of cell passages is that
sufficient evidence must be presented on the chromosomal
number and distribution in each cell line before test data
can be interpreted with confidence. Also, the multiple
scoring of hyper, hypo, and polyploid cells presents a
complex data base, the statistical interpretation of which
might not yet have been optimized. Furthermore, the
possible karyotypic instability of higher passage cells
suggests that the current practice of scoring only 200
cells each in test and control cultures might not be
adequate.
The aneuploidy findings of the CSSTT, therefore, provide an
exciting area for future research. However, the role of such
assays as routinely employed complementary tests must await the
acquisition of further data and the outcome of developmental
studies. The related polyploidy assays are considered in section
9.7.
9.5. Mammalian Cell Gene-Mutation Assays
Mammalian cell gene-mutation assays, together with chromosomal-
aberration assays, are high on the list of optional secondary
assays referred to by legislative authorities. The performance of
these assays was, thus, of particular interest. The mammalian
gene-mutation assays constituted the largest group of assays in the
project.
Sixteen sets of data were considered, 6 of which were based on
nominally the same assay (L5178Y), employing the same selective
agent (trifluorothymidine).
Qualitative inspection of the total data base for the mammalian
gene-mutation assays (Table 2), and of that produced following the
primary reduction step (Table 4), was not, as pointed out by the
working group, initially encouraging. The overall sensitivity of
the collected gene-mutation and chromosomal-aberration assays were
similar (Table 3; 62 and 67% and 46 and 53% overall sensitivity,
respectively, for the 2 groups of 4 carcinogens). The value of
such numerical indices can be questioned, but they are supported by
the individual comparisons drawn in Table 3 for the activity of
each chemical in these 2 classes of assay. Thus, similar overall
sensitivity was observed in both classes of assay for most of the
chemicals. Fig. 4 and 5 show the comparative sensitivity
histograms with the Salmonella assays.
9.3. Assays for DNA Damage SSB (Single-Strand Breaks) and UDS
(Detected via Autoradiography or Scintillation Counting)
Although both of these classes of assay are designed to be
sensitive to the consequences of the primary interaction of the
test chemical (or a metabolite) with DNA, there are good reasons
for not considering them as equivalent. These reasons are
discussed in the assay working group report and are supported by
aspects of the present data base (Tables 2 and 3). For example,
HMPA was inactive in the 3 SSB assays yet was active in the assays
for UDS (2+ and 1?). In contrast, DES was uniformly active in the
3 SSB assays and inactive in the UDS tests. The fact that
unanimity between these 2 assay groups can be achieved is evidenced
by the responses seen for TOL (6/6+), SAF (5/6+), and PB (4/4-).
The relative sensitivity of these 3 classes of assay are shown in
Fig. 3. It should be noted that some legislative authorities have
discussed the need for assay data indicating primary interaction of
a test chemical with DNA. Their requirement could be met with data
generated from either of these groups of assay, i.e., the 2 are
generally regarded as being different methods of assaying a similar
phenomenon. As can be seen, this may not be so.
Perhaps one of the most surprising findings of the CSSTT is the
insensitivity demonstrated by the 2 autoradiographic assays (4.4.1
and 4.4.2, eliminated from Table 4). These were conducted
according to similar protocols and are well represented in earlier
literature. Their poor performance in the present study suggests
that they are unsuitable for use as a complementary assay, though
their use for confirming the activity in mammalian cells of
previously defined bacterial mutagens remains. Another intriguing
aspect of their poor performance is that it provides evidence to
suggest that the autoradiographic end-point is less sensitive than
the other end-points under review (with the exception of the
activity seen for HMPA in one assay, 4.4.1). This insensitivity is
unlikely to be due to metabolic incompetence as the hepatocytes
employed were primary isolates, the metabolic competence of which
had been previously established. Furthermore, the possibility that
this divergence of data was due to these agents producing only
short-patch repair is reduced, given the range of chemically
different carcinogens employed. Whether some of the positive
responses observed in the SSB assays were a direct result of cell
necrosis is discussed below.
Assays for SSB and UDS would be expected to be primarily
sensitive to the 4 confirmed genotoxins represented here, HMPA,
o-toluidine, safrole, and acrylonitrile, and this was observed (cf.
Table 3 and the discussion of the yeast and Drosophila gene-
mutation assays in this section). The only confirmed success with
the remaining 4 carcinogens was the ability of all of the SSB
assays to detect DES. Non-reproducible activity was seen in both
classes of assay for benzene (2/6), benzoin (2/5), and DEHP (1/5).
The latter isolated positive response was strongly endorsed by the
investigator (4.4.3) and requires further investigation.
A topic for future research is the extent to which cell
necrosis, with the consequent formation of broken DNA, contributes
to the outcome of SSB assays. For example, the isolated positive
response observed for benzene (assay 4.3.2) was only observed at
higher dose levels than those employed by the other 2 investigators
in this group. This response may, therefore, represent a technical
false-positive response induced solely by DNA strand breakage as
the result of toxicity. Equally, the extent to which
"mitotic" DNA synthesis has been successfully blocked during the
conduct of a scintillation UDS assay relates directly to the
credibility accorded to weak induced effects. The above 2 concerns
may explain some of the divergencies of data evident within these 3
classes of assay for the present chemicals.
Given the simple requirement to provide evidence of the ability
of a chemical to induce primary DNA damage in mammalian cells, the
investigator is faced with the following decisions:
(a) If the agent is already defined as a bacterial mutagen,
any of the 3 classes of assay (SSB or UDS via
autoradiography or scintillation counting) discussed
herein could be used.
(b) If the agent is inactive as a bacterial mutagen, use of
either the SSB or scintillation counted UDS assays could
present advantages, depending on the chemicals involved.
The results obtained for HMPA and DES suggest that the use
of both assays should be considered, if an assay for "DNA
damage" is required as a complement to bacterial
mutagenicity.
It was, therefore, concluded that the simple legislative
requirement for "evidence of DNA damage" was too vague to be
meaningful. If complementary data are being sought, the
autoradiographic end-point should be avoided and data should be
derived from an SSB assay and/or scintillation counted UDS assay.
The failure of the CSSTT to indicate clearly a single class of
assay, within this group, for routine adoption suggests that
attention should be given to developing the sensitivity of DNA
damage assays to the point that only a single test is required.
The relative performance of these 3 classes of DNA repair assays is
shown in Fig. 3.
9.4. Assays for the Induction of Aneuploidy
In the past, assays for the induction of aneuploidy have not
generally been considered as predictive tests for carcinogenicity.
In the first place, few appropriate in vitro assays existed, and
secondly, the induction of aneuploidy was assumed to have greater
relevance for the promotional, as opposed to the initiatory stages
of carcinogenesis. Thus, while the previous data base has included
the evaluation of a few reference genotoxins such as MNU, it has
mainly focused on cancer-promoting agents such as the phorbol
esters and saccharin, etc. Two investigators have provided the
greater part of the present data base for this class of assay by
presenting data derived from yeast assays (2.4.1.3). In addition,
data derived using mammalian cells were presented (4.9.1) together
with a set of data observed in Aspergillus (2.4.4).
The performance of the aneuploidy assays in the CSSTT was
impressive (Table 2 and 3). In the case of the 2 sets of yeast
data provided from one laboratory (2.4.1 and 2.4.3), a 100%
correlation between the reported carcinogenicity of the test agents
and their activity in vitro was observed. In particular, the
unanimous detection of both DEHP and phenobarbital by these
aneuploidy assays is unique in this study.
The primary conclusion regarding this group of assays is,
therefore, that they present high promise for development as a
class of complementary assays. The development phase should be
influenced by the following considerations, some of which are of
sufficient importance to ensure that they are resolved before any
particular aneuploidy assay is recommended for general adoption.
(a) The findings of the CSSTT suggest that the induction of
aneuploidy may be a critical step in the mechanism of
action of complete carcinogens. These observations
require independent confirmation in other laboratories,
for both the yeast and the mammalian cell systems.
(b) In the case of the D61-M yeast data from the 2
laboratories considered, major discrepancies were evident.
These were all of the same nature and represented the
failure of one laboratory to detect anything beyond
benzene and DES. It is possible that this divergence was
due to technical factors affecting the mitotic or
metabolic activity of the yeast. For example, evidence
from the fungal studies supported an earlier observation
that growing cells are more sensitive to genotoxins than
stationary cells. This could be because of the reduced
metabolic activity of stationary cells. Thus, this
divergence might be due to a relatively minor technical
consideration, but it must be defined, understood,
corrected, and defended in the literature before any
general adoption of yeast aneuploidy assays can be
recommended. The question of whether the D6 strain can be
regarded as repetitive of the D61-M strain also requires
resolution; if it is to be retained, its efficacy must be
confirmed in other laboratories.
(c) The isolated set of Aspergillus data suggests that this
assay is relatively insensitive. Clearly, if the
induction of aneuploidy is to be pursued as a useful
complementary end-point, and if Aspergillus nidulans is to
be selected as the marker organism, a separate data base
will have to be generated to justify this choice.
(d) Requirements similar to those in (c) must apply to the
isolated set of aneuploidy data generated in the Chinese
hamster liver fibroblast assay (4.9.1). This assay is
relatively novel; thus, apart from an expansion of the
data base and the commissioning of independent studies,
the following point requires attention. The cells used in
this study were of between the 9 - 15th passage. The
corollary for using such a spread of cell passages is that
sufficient evidence must be presented on the chromosomal
number and distribution in each cell line before test data
can be interpreted with confidence. Also, the multiple
scoring of hyper, hypo, and polyploid cells presents a
complex data base, the statistical interpretation of which
might not yet have been optimized. Furthermore, the
possible karyotypic instability of higher passage cells
suggests that the current practice of scoring only 200
cells each in test and control cultures might not be
adequate.
The aneuploidy findings of the CSSTT, therefore, provide an
exciting area for future research. However, the role of such
assays as routinely employed complementary tests must await the
acquisition of further data and the outcome of developmental
studies. The related polyploidy assays are considered in section
9.7.
9.5. Mammalian Cell Gene-Mutation Assays
Mammalian cell gene-mutation assays, together with chromosomal-
aberration assays, are high on the list of optional secondary
assays referred to by legislative authorities. The performance of
these assays was, thus, of particular interest. The mammalian
gene-mutation assays constituted the largest group of assays in the
project.
Sixteen sets of data were considered, 6 of which were based on
nominally the same assay (L5178Y), employing the same selective
agent (trifluorothymidine).
Qualitative inspection of the total data base for the mammalian
gene-mutation assays (Table 2), and of that produced following the
primary reduction step (Table 4), was not, as pointed out by the
working group, initially encouraging. The overall sensitivity of
the collected gene-mutation and chromosomal-aberration assays were
similar (Table 3; 62 and 67% and 46 and 53% overall sensitivity,
respectively, for the 2 groups of 4 carcinogens). The value of
such numerical indices can be questioned, but they are supported by
the individual comparisons drawn in Table 3 for the activity of
each chemical in these 2 classes of assay. Thus, similar overall
sensitivity was observed in both classes of assay for most of the
chemicals. Fig. 4 and 5 show the comparative sensitivity
histograms with the Salmonella assays.
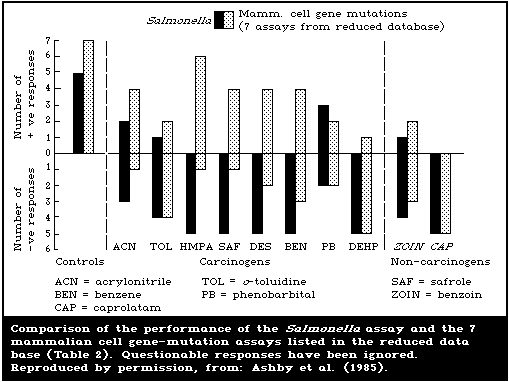
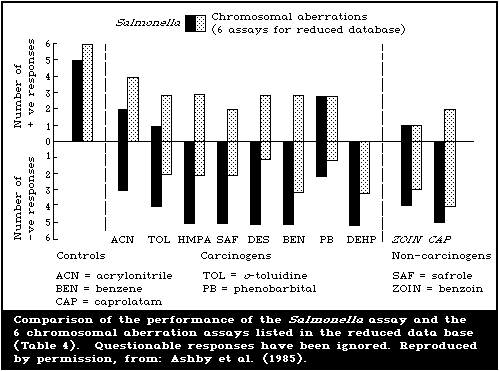 The working-group discussions on the gene-mutation assays
identified a number of specific factors that could have led to the
many disagreements in assay responses and which, if followed up,
should lead to the development of a greatly improved assay system.
Each of these points was concerned with protocol details, all of
which were of relevance to all of these assays, irrespective of the
cell line, selective agent, etc., employed. Some of these
recommendations are of immediate relevance for the interpretation
of weak assay responses (an important matter that is dealt with in
extenso in the full account of the project). The common feature of
the present test chemicals is that they tend to yield weak
responses in vitro. Thus, it is impossible to discern at this
stage which of the many weak positive responses recorded in this
assay group were real and which were illusions created by
inadequate experimentation. Equally, some of the negative
responses observed may have been attributable to experimental
deficiencies rather than to an absolute inadequacy of the assay to
detect the carcinogen. Suggested protocol modifications,
especially those associated with the repetition of experiments and
the use of appropriate cell numbers, should reduce the incidence of
"technical" false-positive and false-negative responses. The data
base that would have been produced in this study, had such
technical modifications already been incorporated, would doubtless
have been much more consistent. However, in addition to possible
protocol deficiencies, which it is anticipated can be remedied,
there remain 2 other important questions.
The first is the extent to which this class of assay can
realistically be reduced to a common cell line using one or more
common selective agents. This would obviously represent a major
advance, as it would enable meaningful interlaboratory comparisons
of data to be made and engender a feeling of confidence that once a
test chemical had been evaluated for mammalian cell gene
mutagenicity, and a conclusion drawn, this would not be reversed on
retesting in another, but similar assay. The need to try to devise
such a system, suitable for general screening was, in fact, the
final recommendation of the assay working group. Some of the
divergencies in test results, evident in this study, were probably
due to the differential sensitivity and metabolic competence of the
several cell lines employed, and this problem cannot be resolved by
adoption of a single common cell line. However, the range of
genetic loci observed and selective agents used probably led to
many instances of true differential sensitivity. The need to use
at least 2 different genetic markers, when a wide range of
genotoxic agents are to be detected has been discussed elsewhere,
and instances of 2 loci being monitored concomitantly have been
reported (Barrett et al., 1981). A considerable amount of
developmental work will be required to discover the degree to which
this class of assay can be standardized with a retention of overall
sensitivity.
The second concern is related to whether these assays are
suitable, by definition, to act as complements to the Salmonella
assays. The results from the yeast and Drosophila assays were not
encouraging and indicated that these particular assays were not
optimal for this purpose, because of the similar nature of the
genetic event being monitored. Similar concerns apply to mammalian
cell gene-mutation assays. Thus, the concept that some agents may
be specifically clastogenic is an argument against the
incorporation of only gene-mutation assays in a battery. Given
that there is usually a requirement in legislative guidelines to
assess a compound for clastogenic activity, the precise role of
mammalian cell gene-mutation assays needs to be reviewed. These
considerations do not necessarily detract from their use as
confirmatory assays. Thus, if it is considered necessary to
evaluate whether a bacterial gene mutagen is similarly active in
mammalian cells, their use is obvious.
The future use of mammalian gene-mutation assays is, therefore,
likely to be influenced by a variety of considerations. The first
and most fundamental is the need for a precise exposition of their
proposed role in the detection of potential carcinogens in vitro.
If they are to be employed only to confirm activity observed in the
Salmonella assay, then some justification should be provided for
the implied existence of a group of established non-carcinogens
that are mutagenic to Salmonella, yet inactive in a well-conducted
mammalian cell gene-mutation assay. If they are to be used as a
complementary assay to the Salmonella assay, urgent attention
should be given to protocol development along the lines outlined in
the assay working group report. The selection of a single assay,
or the development of a single multi-locus assay, may form part of
this development process. However, the wisdom of employing 2 gene-
mutation assays for screening purposes now needs careful
consideration.
Set against these rather demanding requirements are the
following 2 more positive observations. First, it is significant
that mammalian cell gene-mutation assays formed the largest group
of closely-related assays considered in this study. This is
indicative of the current widespread use of these tests and implies
that, in many laboratories, they are performing to an acceptable
standard for reference mutagenic carcinogens such as 2AAF, MNU,
etc. The large number of laboratories carrying out these tests is,
of course, mainly the result of legislative stimulus, but had the
tests proved generally insensitive or unreliable, it would not have
been possible to collect so many together for this study. This is
in contrast to most of the other assays in the study and provides a
strong stimulus for further work to refine them to the point that
they can be used with confidence for the detection of weak
responses. Second, although this study was concerned primarily
with the detection in vitro of potential mammalian carcinogens, the
possible mutagenic properties of chemicals for in vivo somatic and
germ cells are becoming of increasing interest. If the specific
requirement to assay for possible mammalian in vivo mutagens is a
consideration, the role of in vitro gene-mutation assays assumes a
different complexion.
9.6. Chromosomal-Aberration Assays
As indicated above, the performance of this class of assay was
similar to, but slightly better than, that of the mammalian cell
gene-mutation assays. As with the gene-mutation assays,
chromosomal-aberration tests are generally recommended in almost
all legislative guidelines as suitable for use in conjunction with
a bacterial-mutation test, when screening chemicals for new
potential carcinogens. However, while it is currently normal
practice to select the L5178Y cell line when conducting a gene-
mutation experiment, a range of possible cell lines could be chosen
for cytogenetic studies. Thus, in the CSSTT, 4 distinct cell lines
(CHO cells, human lymphocytes, Chinese hamster liver fibroblasts
(CHI-L), and Chinese hamster lung cells (CHL)) were selected, each
of which had been used previously for such studies, and each of
which could be considered suitable for use, when submitting
genotoxicity test data on a new chemical.
A distinct attribute of the cytogenetics assay data was that,
although a range of cytogenetic disturbances can be classified as
chemically-induced damage, the data base was more ordered than
might have been anticipated, because of the common reporting format
employed by most investigators. This was achieved, in advance, by
the assay group Chairman and was based substantially on the
guidelines for assay data reporting suggested in the UKEMS criteria
document (UKEMS, 1983). Further sources of protocol variation can,
therefore, be associated directly with technical decisions made by
individual investigators, including such issues as doselevels,
sampling times, methods of statistical analysis, cell line adopted,
etc.
As would be expected from the similar overall detection rates
of the gene-mutation and chromosomal-aberration assays (Table 3,
and Fig. 4 and 5), simple inspection of the chromosomal-aberration
data base (Table 2 and 4) indicates a disappointing scatter of
positive and negative observations. Certain consistencies were,
however, evident. For example, general agreement was observed in
the 3 cell lines studied for the clastogenicity of both
acrylonitrile (4/4+) and phenobarbital (3+ and 1? of 4). With the
exception of a questionable response in one of the 3 CHO assays
(4.5.3), DEHP gave negative results. Benzoin gave 3 negative
responses and a single, but confirmed, positive response in the CHL
assay. A partially consistent data set was observed for HMPA,
which gave a negative response in each of the 2 CHO assays but was
active in the 3 other cytogenetic assays. This may reflect the
general technical inadequacy of these assay protocols or represent
a general failure of this cell line to metabolize HMPA, a genotoxic
species. One of the 2 CHO SCE assays gave a positive response for
this agent, and this weakens, but does not destroy, the metabolic
deficiency rationale.
Marked divergencies in assay responses were evident for the
remaining 5 chemicals ( o-toluidine, safrole, benzene, caprolactam,
and DES). An approximately equal incidence of positive and
negative responses was observed for these agents and each was
regarded as clastogenic in the assay working-group "consensus"
results ( o-toluidine 3/5 + safrole 2/4 +, benzene 3/6 + caprolactam
2/6 +, and DES 3/4 +). Important protocol deficiencies were
identified in the assay working-group discussions. These
deficiencies were generally associated with the selection of
inappropriate sampling times or dose levels, and if further work
could prove them to be the critical contributors to false-negative
assay responses, then the performance of these assays would be
dramatically improved.
For the purpose of defining a complementary assay within the
present context, the 4 classes of chromosomal-aberration assays
cannot be considered as equivalent. For example, the CHL assay has
been extensively studied and reported on in the literature, but its
use is centred mainly in a single laboratory. Assays involving
human lymphocytes, especially if a rat liver S9 mix is incorporated
into the protocol, are few (Buckton & Evans, 1982). The study data
using Chinese hamster liver fibroblasts formed the greater part of
the accepted data base for this assay. In contrast, the CHO assay
probably represented the one most widely used in industrial and
academic laboratories. This assay can, therefore, be regarded as
particularly relevant for the objectives of the project, as
probably more data are generated using it than any other, when
compounds are assessed prior to legal registration. Given that
some of the negative responses observed in CHO cells might be
corrected by the suggested modifications to the assay protocol,
there is still the problem that, at present, different responses
may be observed in different laboratories using nominally the same
assay. This is similar to the situation currently evident with the
L5178Y gene-mutation assays and, as in that case, urgent attention
to further protocol design and data interpretation is indicated.
The CHL assay (4.5.6), or perhaps the protocol by which it was
conducted, was more sensitive than the others in this group. By
the same criteria, the partial data base evident for the human
lymphocyte assay was also encouraging and worthy of elaboration.
As in other assay systems, it was noted how "assays" were judged,
or misjudged, by the test protocols adopted. This important aspect
of test systems must be resolved, because of the real possibility
that, if more detailed or refined protocols had been used, many
assays would have appeared in a much more favourable light.
An advantage of the chromosomal assays is the extent of the
existing data base and the technical skill that is available for
the conduct of these assays. Thus, it can be expected that
acceptable protocol modifications could be rapidly developed and
disseminated for general adoption. A second advantage is that a
range of supplementary and probably "complementary" end-points can
be easily assessed, once the decision has been taken to conduct
cytogenetic studies within a laboratory. These include the
recording of polyploid cells, an assessment of aneuploidy, and the
studying of sister chromatid exchanges (SCE). Aneuploidy induction
in mammalian cells is considered as a separate assay, and the
induction of SCEs is discussed below. However, the induction of
polyploidy is pertinent to the present discussion as it can be
assessed either concomitantly with, or in parallel to, chromosomal-
aberration assays, depending on the sampling time of the assay.
9.7. Assays for Polyploidy Induction
The chemical induction of polyploidy was discounted as a
separate assay by the working group, but the data generated were
considered because of the interest of the results that were
available. Three complete sets of polyploidy data were presented
(Table 2; assays 4.8.1 (CHL), 4.8.2 (CH liver), and 4.8.3 (RL4, rat
liver)), and each detected DES as positive. This activity is
probably associated with the ability of DES to induce aneuploidy,
spindle effects, and chromosomal aberrations. In the last 2 assays
(4.8.2 and 4.8.3), the remaining 9 compounds were found to be
negative while, in the CHL assay (4.8.1), o-toluidine and
caprolactam were detected as active. These responses indicate that
this genetic end-point is unlikely to produce false-positive
responses; in fact, activity seems to be rare.
The mode of action of DES, benzene, and DEHP as carcinogens is
less clearly defined than that of most other agents. Not only are
they inactive in the Salmonella assay, but they were difficult to
detect as positive in many of the eukaryotic assays used in the
CSSTT. Furthermore, on the basis of their chemical structure and
known or anticipated metabolism, these agents do not appear to be
likely to interact covalently with DNA. It was, therefore,
perceived by many investigators to be of particular interest that
each of these agents was capable of inducing both aneuploidy and
cell transformation in vitro. Furthermore, if "associated"
chromosomal-assay data are considered in conjunction with the
aneuploidy findings, a potentially useful pattern emerges, as shown
in Table 5.
Table 5. Comparative performance of DES, benzene, and DEHP in the classes of
assay showna
----------------------------------------------------------------------------------
Cell Aneuploidy Clastogenicity Polyploidy Spindle studies
transformation (assay 4.9.2)
----------------------------------------------------------------------------------
DES + + + + spindle damage
benzene + + + - modified spindle
function
DEHP + + - - modified spindle
function
----------------------------------------------------------------------------------
a Results are based on the consensus conclusions given by the working groups.
From Table 5, it can be seen that DES was clearly active in
each of the 3 categories of assay, and that its mode of action in
each might be closely related to its ability to damage the
microtubules of the metaphase spindle. In contrast, DEHP was
inactive in the 3 polyploidy assays, as well as in the chromosomal
assays. The question therefore arises of whether the cell-
transforming and aneuploidy-inducing properties of DEHP are
mechanistically unrelated to those of DES. If totally different
mechanisms of action are involved, then the relationship of these
findings to the carcinogenicity of these agents may equally be
different. In this connection, the difference between modification
of spindle integrity and spindle function may be important. Thus,
it is concluded that if an assay for chromosomal aberrations is to
be used as a complementary test, the determination of changes in
ploidy may be a useful additional parameter to measure. In some
assays, it may not be possible to score both chromosomal
aberrations and ploidy, at the same time, mainly because of the
need to progress to the second mitosis after treatment, for
polyploidy to become evident. However, the maintenance of a
parallel culture after treatment for separate assessment of ploidy
should present few difficulties.
9.8. Sister Chromatid Exchange (SCE) Assays
The relationship between the chemical induction of SCEs and the
production of chromosomal aberrations is not clear, and the
mechanism of this phenomenon is obscure. The role of SCE assays in
the detection of carcinogens has yet to be defined. For these
reasons, and because their respective test protocols are quite
distinct, SCE assays have been considered separately from the
aberration assays in the present discussion.
One set of data was omitted from Table 4 owing to the recording
of 6 out of 6 negative responses (4.6.4) and, in the remaining 5
SCE assays, only 12 positive responses were observed out of 41
determinations. These positive responses were not randomly
distributed among the compounds as can be seen in Table 3. Two-
thirds of the SCE assays made on HMPA, o-toluidine, safrole, and
acrylonitrile gave positive responses, while only a single positive
response was observed among the 14 determinations made on benzene,
DEHP, DES, or phenobarbital. This grouping of positive responses
is very similar to that seen with the Drosophila mutation assays
and to a lesser extent by the yeast gene-mutation and UDS
(scintillation) assays. This supports earlier suggestions that the
induction of SCEs is more closely related to gene mutagenicity than
to clastogenicity. No activity was observed for the 2 non-
carcinogens.
Given that the greatest number of positive responses was
observed in this study for the first 4 "genotoxic" carcinogens
shown in Table 4, the SCE findings are unexceptional. The
inability to detect the remaining 4 carcinogens by SCE assays may
provide clues (along with the Drosophila assays) regarding the
mechanism of carcinogenic action of these agents. An observation
of particular interest is that 3 carcinogens, benzene, DES, and
phenobarbital (but not DEHP) were each active in one or more of the
other chromosome-based assays; their inactivity in the SCE assays,
therefore, becomes pointed.
The SCE assays, therefore, appear to be of limited advantage as
complementary assays. Together with several other assays, they
successfully detected the "agreed" genotoxins in the CSSTT and, as
such, the assay seems to be more closely related to the Drosophila
and yeast gene-mutation assays. The low detection rates evident
for the 4 remaining carcinogens suggest that SCE assays are less
generally useful as a complementary test than the chromosomal-
aberration and mammalian cell gene-mutation assays.
9.9. Transformation Assays
The working group, for reasons briefly referred to earlier, did
not consider this class of assay to be suitable for a generally-
applicable complementary assay. However, the good overall
performance of the assay in the CSSTT (Table 3) warrants further
consideration. The chemical induction, in vitro, of the
transformed cell phenotype is appealing, because the derived cell
morphology and behaviour in vivo and in vitro bear a striking
resemblance to the malignant phenotype. The many different
definitions of the transformed cell phenotype (Weinstein et al.,
1976), together with the technical difficulties that accompany the
conduct of such assays have proved to be major factors contributing
to the slow progress that has been made in the general use of these
tests for the detection of possible new carcinogens. The initial
promise shown by the BHK-21 cell-transformation assay was due to
the compatibility of these cells with S9 mix (Styles, 1977), but
progress was impeded by the karyotypic instability of this cell
line and inter-laboratory reproducibility problems encountered at
an early stage (de Serres & Ashby, 1981). The present study
represents the first occasion on which the mouse C3H 10T 1/2 assay
and the Syrian hamster embryo (SHE) assay have been directly
compared in a general study, yet each has an impressive history in
a few laboratories. Although both of these classes of assay
performed well in the CSSTT, there was a marked reluctance among
the assay investigators when it came to suggesting any of these
tests as suitable for routine adoption as a complementary assay.
The insensitivity to the present carcinogens of the Balb C assay
(4.2.1) and the CHO assay (4.2.8) confirmed previous observations
that established cell lines tend to become metabolically
incompetent, yet the metabolically competent primary cell systems
suffer from other problems associated with the criteria used to
select clones of cells for study, and the selection of appropriate
media.
Recent studies associated with the probable role of activation
of the oncogene in the etiology of cancer have invoked the
transformed cell phenotype as a critical marker; thus, the future
of this class of assay in cancer research seems assured.
Nonetheless, the practical problems that would probably be
encountered by a new laboratory endeavouring to establish such an
assay are probably still sufficient to warn against its use as a
routinely-employed complementary test.
10. SELECTION OF A PREFERRED COMPLEMENTARY ASSAY
No single assay was defined by the CSSTT as giving high
carcinogen sensitivity and specificity together with an established
ability to perform optimally in several laboratories. It was
concluded, however, that there was compelling evidence both from
within and beyond the collaborative study, for the adoption of a
chromosomal-aberration assay as the reference complementary test.
This conclusion was drawn on the basis of the following data and
reasoning:
(a) Eight classes of tests, as listed in Table 3, were assessed
for performance as complementary assays, in the foregoing
discussion. Of these, the gene-mutation assays in yeast
and Drosophila were not favoured because of their
repetition of genetic end-point and their poor performance
with the last 4 carcinogens (benzene, DES, phenobarbital,
and DEHP). Assays for aneuploidy were not recommended
because of their novelty, which necessarily prevented
independent studies from being commissioned. The DNA-
repair assays showed poor sensitivity for the last 4
carcinogens, and, occasionally, dramatic differences in
sensitivity between the single-strand breakage and
scintillation UDS tests were evident (Fig. 3). The data
base for the SCE assays suggested marked insensitivity to
the last 4 carcinogens (Fig. 6). Several of the above
assays were, therefore, defined as capable of acting as a
partial complement to the Salmonella assay, but none could
be recommended for routine use as a generally sensitive
complementary test.
The working-group discussions on the gene-mutation assays
identified a number of specific factors that could have led to the
many disagreements in assay responses and which, if followed up,
should lead to the development of a greatly improved assay system.
Each of these points was concerned with protocol details, all of
which were of relevance to all of these assays, irrespective of the
cell line, selective agent, etc., employed. Some of these
recommendations are of immediate relevance for the interpretation
of weak assay responses (an important matter that is dealt with in
extenso in the full account of the project). The common feature of
the present test chemicals is that they tend to yield weak
responses in vitro. Thus, it is impossible to discern at this
stage which of the many weak positive responses recorded in this
assay group were real and which were illusions created by
inadequate experimentation. Equally, some of the negative
responses observed may have been attributable to experimental
deficiencies rather than to an absolute inadequacy of the assay to
detect the carcinogen. Suggested protocol modifications,
especially those associated with the repetition of experiments and
the use of appropriate cell numbers, should reduce the incidence of
"technical" false-positive and false-negative responses. The data
base that would have been produced in this study, had such
technical modifications already been incorporated, would doubtless
have been much more consistent. However, in addition to possible
protocol deficiencies, which it is anticipated can be remedied,
there remain 2 other important questions.
The first is the extent to which this class of assay can
realistically be reduced to a common cell line using one or more
common selective agents. This would obviously represent a major
advance, as it would enable meaningful interlaboratory comparisons
of data to be made and engender a feeling of confidence that once a
test chemical had been evaluated for mammalian cell gene
mutagenicity, and a conclusion drawn, this would not be reversed on
retesting in another, but similar assay. The need to try to devise
such a system, suitable for general screening was, in fact, the
final recommendation of the assay working group. Some of the
divergencies in test results, evident in this study, were probably
due to the differential sensitivity and metabolic competence of the
several cell lines employed, and this problem cannot be resolved by
adoption of a single common cell line. However, the range of
genetic loci observed and selective agents used probably led to
many instances of true differential sensitivity. The need to use
at least 2 different genetic markers, when a wide range of
genotoxic agents are to be detected has been discussed elsewhere,
and instances of 2 loci being monitored concomitantly have been
reported (Barrett et al., 1981). A considerable amount of
developmental work will be required to discover the degree to which
this class of assay can be standardized with a retention of overall
sensitivity.
The second concern is related to whether these assays are
suitable, by definition, to act as complements to the Salmonella
assays. The results from the yeast and Drosophila assays were not
encouraging and indicated that these particular assays were not
optimal for this purpose, because of the similar nature of the
genetic event being monitored. Similar concerns apply to mammalian
cell gene-mutation assays. Thus, the concept that some agents may
be specifically clastogenic is an argument against the
incorporation of only gene-mutation assays in a battery. Given
that there is usually a requirement in legislative guidelines to
assess a compound for clastogenic activity, the precise role of
mammalian cell gene-mutation assays needs to be reviewed. These
considerations do not necessarily detract from their use as
confirmatory assays. Thus, if it is considered necessary to
evaluate whether a bacterial gene mutagen is similarly active in
mammalian cells, their use is obvious.
The future use of mammalian gene-mutation assays is, therefore,
likely to be influenced by a variety of considerations. The first
and most fundamental is the need for a precise exposition of their
proposed role in the detection of potential carcinogens in vitro.
If they are to be employed only to confirm activity observed in the
Salmonella assay, then some justification should be provided for
the implied existence of a group of established non-carcinogens
that are mutagenic to Salmonella, yet inactive in a well-conducted
mammalian cell gene-mutation assay. If they are to be used as a
complementary assay to the Salmonella assay, urgent attention
should be given to protocol development along the lines outlined in
the assay working group report. The selection of a single assay,
or the development of a single multi-locus assay, may form part of
this development process. However, the wisdom of employing 2 gene-
mutation assays for screening purposes now needs careful
consideration.
Set against these rather demanding requirements are the
following 2 more positive observations. First, it is significant
that mammalian cell gene-mutation assays formed the largest group
of closely-related assays considered in this study. This is
indicative of the current widespread use of these tests and implies
that, in many laboratories, they are performing to an acceptable
standard for reference mutagenic carcinogens such as 2AAF, MNU,
etc. The large number of laboratories carrying out these tests is,
of course, mainly the result of legislative stimulus, but had the
tests proved generally insensitive or unreliable, it would not have
been possible to collect so many together for this study. This is
in contrast to most of the other assays in the study and provides a
strong stimulus for further work to refine them to the point that
they can be used with confidence for the detection of weak
responses. Second, although this study was concerned primarily
with the detection in vitro of potential mammalian carcinogens, the
possible mutagenic properties of chemicals for in vivo somatic and
germ cells are becoming of increasing interest. If the specific
requirement to assay for possible mammalian in vivo mutagens is a
consideration, the role of in vitro gene-mutation assays assumes a
different complexion.
9.6. Chromosomal-Aberration Assays
As indicated above, the performance of this class of assay was
similar to, but slightly better than, that of the mammalian cell
gene-mutation assays. As with the gene-mutation assays,
chromosomal-aberration tests are generally recommended in almost
all legislative guidelines as suitable for use in conjunction with
a bacterial-mutation test, when screening chemicals for new
potential carcinogens. However, while it is currently normal
practice to select the L5178Y cell line when conducting a gene-
mutation experiment, a range of possible cell lines could be chosen
for cytogenetic studies. Thus, in the CSSTT, 4 distinct cell lines
(CHO cells, human lymphocytes, Chinese hamster liver fibroblasts
(CHI-L), and Chinese hamster lung cells (CHL)) were selected, each
of which had been used previously for such studies, and each of
which could be considered suitable for use, when submitting
genotoxicity test data on a new chemical.
A distinct attribute of the cytogenetics assay data was that,
although a range of cytogenetic disturbances can be classified as
chemically-induced damage, the data base was more ordered than
might have been anticipated, because of the common reporting format
employed by most investigators. This was achieved, in advance, by
the assay group Chairman and was based substantially on the
guidelines for assay data reporting suggested in the UKEMS criteria
document (UKEMS, 1983). Further sources of protocol variation can,
therefore, be associated directly with technical decisions made by
individual investigators, including such issues as doselevels,
sampling times, methods of statistical analysis, cell line adopted,
etc.
As would be expected from the similar overall detection rates
of the gene-mutation and chromosomal-aberration assays (Table 3,
and Fig. 4 and 5), simple inspection of the chromosomal-aberration
data base (Table 2 and 4) indicates a disappointing scatter of
positive and negative observations. Certain consistencies were,
however, evident. For example, general agreement was observed in
the 3 cell lines studied for the clastogenicity of both
acrylonitrile (4/4+) and phenobarbital (3+ and 1? of 4). With the
exception of a questionable response in one of the 3 CHO assays
(4.5.3), DEHP gave negative results. Benzoin gave 3 negative
responses and a single, but confirmed, positive response in the CHL
assay. A partially consistent data set was observed for HMPA,
which gave a negative response in each of the 2 CHO assays but was
active in the 3 other cytogenetic assays. This may reflect the
general technical inadequacy of these assay protocols or represent
a general failure of this cell line to metabolize HMPA, a genotoxic
species. One of the 2 CHO SCE assays gave a positive response for
this agent, and this weakens, but does not destroy, the metabolic
deficiency rationale.
Marked divergencies in assay responses were evident for the
remaining 5 chemicals ( o-toluidine, safrole, benzene, caprolactam,
and DES). An approximately equal incidence of positive and
negative responses was observed for these agents and each was
regarded as clastogenic in the assay working-group "consensus"
results ( o-toluidine 3/5 + safrole 2/4 +, benzene 3/6 + caprolactam
2/6 +, and DES 3/4 +). Important protocol deficiencies were
identified in the assay working-group discussions. These
deficiencies were generally associated with the selection of
inappropriate sampling times or dose levels, and if further work
could prove them to be the critical contributors to false-negative
assay responses, then the performance of these assays would be
dramatically improved.
For the purpose of defining a complementary assay within the
present context, the 4 classes of chromosomal-aberration assays
cannot be considered as equivalent. For example, the CHL assay has
been extensively studied and reported on in the literature, but its
use is centred mainly in a single laboratory. Assays involving
human lymphocytes, especially if a rat liver S9 mix is incorporated
into the protocol, are few (Buckton & Evans, 1982). The study data
using Chinese hamster liver fibroblasts formed the greater part of
the accepted data base for this assay. In contrast, the CHO assay
probably represented the one most widely used in industrial and
academic laboratories. This assay can, therefore, be regarded as
particularly relevant for the objectives of the project, as
probably more data are generated using it than any other, when
compounds are assessed prior to legal registration. Given that
some of the negative responses observed in CHO cells might be
corrected by the suggested modifications to the assay protocol,
there is still the problem that, at present, different responses
may be observed in different laboratories using nominally the same
assay. This is similar to the situation currently evident with the
L5178Y gene-mutation assays and, as in that case, urgent attention
to further protocol design and data interpretation is indicated.
The CHL assay (4.5.6), or perhaps the protocol by which it was
conducted, was more sensitive than the others in this group. By
the same criteria, the partial data base evident for the human
lymphocyte assay was also encouraging and worthy of elaboration.
As in other assay systems, it was noted how "assays" were judged,
or misjudged, by the test protocols adopted. This important aspect
of test systems must be resolved, because of the real possibility
that, if more detailed or refined protocols had been used, many
assays would have appeared in a much more favourable light.
An advantage of the chromosomal assays is the extent of the
existing data base and the technical skill that is available for
the conduct of these assays. Thus, it can be expected that
acceptable protocol modifications could be rapidly developed and
disseminated for general adoption. A second advantage is that a
range of supplementary and probably "complementary" end-points can
be easily assessed, once the decision has been taken to conduct
cytogenetic studies within a laboratory. These include the
recording of polyploid cells, an assessment of aneuploidy, and the
studying of sister chromatid exchanges (SCE). Aneuploidy induction
in mammalian cells is considered as a separate assay, and the
induction of SCEs is discussed below. However, the induction of
polyploidy is pertinent to the present discussion as it can be
assessed either concomitantly with, or in parallel to, chromosomal-
aberration assays, depending on the sampling time of the assay.
9.7. Assays for Polyploidy Induction
The chemical induction of polyploidy was discounted as a
separate assay by the working group, but the data generated were
considered because of the interest of the results that were
available. Three complete sets of polyploidy data were presented
(Table 2; assays 4.8.1 (CHL), 4.8.2 (CH liver), and 4.8.3 (RL4, rat
liver)), and each detected DES as positive. This activity is
probably associated with the ability of DES to induce aneuploidy,
spindle effects, and chromosomal aberrations. In the last 2 assays
(4.8.2 and 4.8.3), the remaining 9 compounds were found to be
negative while, in the CHL assay (4.8.1), o-toluidine and
caprolactam were detected as active. These responses indicate that
this genetic end-point is unlikely to produce false-positive
responses; in fact, activity seems to be rare.
The mode of action of DES, benzene, and DEHP as carcinogens is
less clearly defined than that of most other agents. Not only are
they inactive in the Salmonella assay, but they were difficult to
detect as positive in many of the eukaryotic assays used in the
CSSTT. Furthermore, on the basis of their chemical structure and
known or anticipated metabolism, these agents do not appear to be
likely to interact covalently with DNA. It was, therefore,
perceived by many investigators to be of particular interest that
each of these agents was capable of inducing both aneuploidy and
cell transformation in vitro. Furthermore, if "associated"
chromosomal-assay data are considered in conjunction with the
aneuploidy findings, a potentially useful pattern emerges, as shown
in Table 5.
Table 5. Comparative performance of DES, benzene, and DEHP in the classes of
assay showna
----------------------------------------------------------------------------------
Cell Aneuploidy Clastogenicity Polyploidy Spindle studies
transformation (assay 4.9.2)
----------------------------------------------------------------------------------
DES + + + + spindle damage
benzene + + + - modified spindle
function
DEHP + + - - modified spindle
function
----------------------------------------------------------------------------------
a Results are based on the consensus conclusions given by the working groups.
From Table 5, it can be seen that DES was clearly active in
each of the 3 categories of assay, and that its mode of action in
each might be closely related to its ability to damage the
microtubules of the metaphase spindle. In contrast, DEHP was
inactive in the 3 polyploidy assays, as well as in the chromosomal
assays. The question therefore arises of whether the cell-
transforming and aneuploidy-inducing properties of DEHP are
mechanistically unrelated to those of DES. If totally different
mechanisms of action are involved, then the relationship of these
findings to the carcinogenicity of these agents may equally be
different. In this connection, the difference between modification
of spindle integrity and spindle function may be important. Thus,
it is concluded that if an assay for chromosomal aberrations is to
be used as a complementary test, the determination of changes in
ploidy may be a useful additional parameter to measure. In some
assays, it may not be possible to score both chromosomal
aberrations and ploidy, at the same time, mainly because of the
need to progress to the second mitosis after treatment, for
polyploidy to become evident. However, the maintenance of a
parallel culture after treatment for separate assessment of ploidy
should present few difficulties.
9.8. Sister Chromatid Exchange (SCE) Assays
The relationship between the chemical induction of SCEs and the
production of chromosomal aberrations is not clear, and the
mechanism of this phenomenon is obscure. The role of SCE assays in
the detection of carcinogens has yet to be defined. For these
reasons, and because their respective test protocols are quite
distinct, SCE assays have been considered separately from the
aberration assays in the present discussion.
One set of data was omitted from Table 4 owing to the recording
of 6 out of 6 negative responses (4.6.4) and, in the remaining 5
SCE assays, only 12 positive responses were observed out of 41
determinations. These positive responses were not randomly
distributed among the compounds as can be seen in Table 3. Two-
thirds of the SCE assays made on HMPA, o-toluidine, safrole, and
acrylonitrile gave positive responses, while only a single positive
response was observed among the 14 determinations made on benzene,
DEHP, DES, or phenobarbital. This grouping of positive responses
is very similar to that seen with the Drosophila mutation assays
and to a lesser extent by the yeast gene-mutation and UDS
(scintillation) assays. This supports earlier suggestions that the
induction of SCEs is more closely related to gene mutagenicity than
to clastogenicity. No activity was observed for the 2 non-
carcinogens.
Given that the greatest number of positive responses was
observed in this study for the first 4 "genotoxic" carcinogens
shown in Table 4, the SCE findings are unexceptional. The
inability to detect the remaining 4 carcinogens by SCE assays may
provide clues (along with the Drosophila assays) regarding the
mechanism of carcinogenic action of these agents. An observation
of particular interest is that 3 carcinogens, benzene, DES, and
phenobarbital (but not DEHP) were each active in one or more of the
other chromosome-based assays; their inactivity in the SCE assays,
therefore, becomes pointed.
The SCE assays, therefore, appear to be of limited advantage as
complementary assays. Together with several other assays, they
successfully detected the "agreed" genotoxins in the CSSTT and, as
such, the assay seems to be more closely related to the Drosophila
and yeast gene-mutation assays. The low detection rates evident
for the 4 remaining carcinogens suggest that SCE assays are less
generally useful as a complementary test than the chromosomal-
aberration and mammalian cell gene-mutation assays.
9.9. Transformation Assays
The working group, for reasons briefly referred to earlier, did
not consider this class of assay to be suitable for a generally-
applicable complementary assay. However, the good overall
performance of the assay in the CSSTT (Table 3) warrants further
consideration. The chemical induction, in vitro, of the
transformed cell phenotype is appealing, because the derived cell
morphology and behaviour in vivo and in vitro bear a striking
resemblance to the malignant phenotype. The many different
definitions of the transformed cell phenotype (Weinstein et al.,
1976), together with the technical difficulties that accompany the
conduct of such assays have proved to be major factors contributing
to the slow progress that has been made in the general use of these
tests for the detection of possible new carcinogens. The initial
promise shown by the BHK-21 cell-transformation assay was due to
the compatibility of these cells with S9 mix (Styles, 1977), but
progress was impeded by the karyotypic instability of this cell
line and inter-laboratory reproducibility problems encountered at
an early stage (de Serres & Ashby, 1981). The present study
represents the first occasion on which the mouse C3H 10T 1/2 assay
and the Syrian hamster embryo (SHE) assay have been directly
compared in a general study, yet each has an impressive history in
a few laboratories. Although both of these classes of assay
performed well in the CSSTT, there was a marked reluctance among
the assay investigators when it came to suggesting any of these
tests as suitable for routine adoption as a complementary assay.
The insensitivity to the present carcinogens of the Balb C assay
(4.2.1) and the CHO assay (4.2.8) confirmed previous observations
that established cell lines tend to become metabolically
incompetent, yet the metabolically competent primary cell systems
suffer from other problems associated with the criteria used to
select clones of cells for study, and the selection of appropriate
media.
Recent studies associated with the probable role of activation
of the oncogene in the etiology of cancer have invoked the
transformed cell phenotype as a critical marker; thus, the future
of this class of assay in cancer research seems assured.
Nonetheless, the practical problems that would probably be
encountered by a new laboratory endeavouring to establish such an
assay are probably still sufficient to warn against its use as a
routinely-employed complementary test.
10. SELECTION OF A PREFERRED COMPLEMENTARY ASSAY
No single assay was defined by the CSSTT as giving high
carcinogen sensitivity and specificity together with an established
ability to perform optimally in several laboratories. It was
concluded, however, that there was compelling evidence both from
within and beyond the collaborative study, for the adoption of a
chromosomal-aberration assay as the reference complementary test.
This conclusion was drawn on the basis of the following data and
reasoning:
(a) Eight classes of tests, as listed in Table 3, were assessed
for performance as complementary assays, in the foregoing
discussion. Of these, the gene-mutation assays in yeast
and Drosophila were not favoured because of their
repetition of genetic end-point and their poor performance
with the last 4 carcinogens (benzene, DES, phenobarbital,
and DEHP). Assays for aneuploidy were not recommended
because of their novelty, which necessarily prevented
independent studies from being commissioned. The DNA-
repair assays showed poor sensitivity for the last 4
carcinogens, and, occasionally, dramatic differences in
sensitivity between the single-strand breakage and
scintillation UDS tests were evident (Fig. 3). The data
base for the SCE assays suggested marked insensitivity to
the last 4 carcinogens (Fig. 6). Several of the above
assays were, therefore, defined as capable of acting as a
partial complement to the Salmonella assay, but none could
be recommended for routine use as a generally sensitive
complementary test.
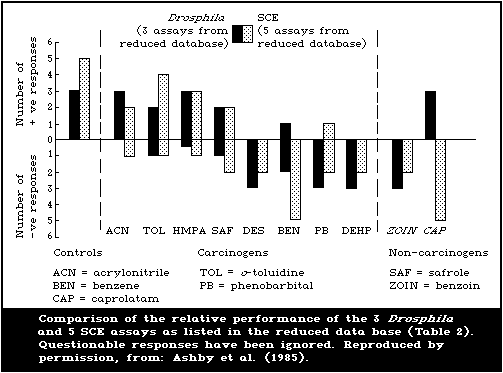 (b) The transformation working group recommended that assays
for cell transformation were not suitable for routine
adoption because of the technical difficulties of carrying
them out and the remaining uncertainty regarding the
definition of the transformed phenotype. Nonetheless, 6 of
these assays showed high sensitivity to the present 8
carcinogens; in fact, as a class, they performed better
than any other category of test (Table 3). It is clear
that both the C3H 10T 1/2 and the SHE assay are sensitive
assays, and each performed similarly, in the CSSTT, in 2
different laboratories. Nonetheless, the advice of the
assay group is endorsed here; namely, that transformation
assays are not to be recommended for routine adoption, at
present, especially by laboratories new to such techniques.
(c) Two classes of assay remain for consideration, each of
which was well represented in the CSSTT, i.e., mammalian
cell gene-mutation and chromosomal-aberration assays. Both
of these classes of assay showed a similar level of
carcinogen sensitivity, when considering the reduced data
base (Table 3 and 4).
The apparently similar performance of these 2 classes of
assay as complementary tests is shown in Fig. 4 and 5. The
performance of the Drosophila and SCE assays is shown in
Fig. 6 for purposes of comparison and to illustrate that
some assays only partially complement the Salmonella assay.
When considering the total data base (Table 2) for the
gene-mutation and chromosomal-aberration assays, the latter
appear to be more sensitive, yielding 22 positive responses
out of 43 determinations made on the 8 carcinogens (51%),
as opposed to only 40 out of 103 for the mutation assays
(39%). The low sensitivity of some of the mutation assays
led to 6 of the 16 being eliminated in Table 3 as opposed
to only 1 of the 7 cytogenetic assays. The normalized (%)
performances of these 2 classes of assays as complementary
tests are compared in Fig. 7. Similar efficiencies are
evident with the chromosomal assays proving marginally more
sensitive to the carcinogens.
Figs. 4, 5, and 7 enable the problem of reproducibility of
response to be visualized and it is clear that harmonization and/or
extension of assay protocols is indicated for both classes of test.
The effort required to produce a common and sensitive assay
protocol, combined with a realistic assessment of whether such
efforts are likely to be rewarded, constitutes the final factor in
the recommendation of the chromosomal-aberration assays. Four
points appear to be particularly relevant:
(a) chromosomal-aberration assays offer an independent genetic
end-point from the Salmonella assay;
(b) chromosomal-aberration assays have a more extensive
history than the mammalian cell gene-mutation assays; this
means that more laboratories have used them for a longer
period; thus, any protocol corrections required should be
more clearly perceived and easier to institute;
(c) the recommendations for assay protocol modification made
by the chromosomal-aberration working group were more
clearly defined, less fundamental, and seemingly easier to
institute than those made by the gene-mutation working
group; and
(d) once the decision is made to conduct a chromosomal-
aberration assay in a laboratory, the expertise and
facilities required will serve equally for the assessment
of aneuploidy, polyploidy, and SCEs.
(b) The transformation working group recommended that assays
for cell transformation were not suitable for routine
adoption because of the technical difficulties of carrying
them out and the remaining uncertainty regarding the
definition of the transformed phenotype. Nonetheless, 6 of
these assays showed high sensitivity to the present 8
carcinogens; in fact, as a class, they performed better
than any other category of test (Table 3). It is clear
that both the C3H 10T 1/2 and the SHE assay are sensitive
assays, and each performed similarly, in the CSSTT, in 2
different laboratories. Nonetheless, the advice of the
assay group is endorsed here; namely, that transformation
assays are not to be recommended for routine adoption, at
present, especially by laboratories new to such techniques.
(c) Two classes of assay remain for consideration, each of
which was well represented in the CSSTT, i.e., mammalian
cell gene-mutation and chromosomal-aberration assays. Both
of these classes of assay showed a similar level of
carcinogen sensitivity, when considering the reduced data
base (Table 3 and 4).
The apparently similar performance of these 2 classes of
assay as complementary tests is shown in Fig. 4 and 5. The
performance of the Drosophila and SCE assays is shown in
Fig. 6 for purposes of comparison and to illustrate that
some assays only partially complement the Salmonella assay.
When considering the total data base (Table 2) for the
gene-mutation and chromosomal-aberration assays, the latter
appear to be more sensitive, yielding 22 positive responses
out of 43 determinations made on the 8 carcinogens (51%),
as opposed to only 40 out of 103 for the mutation assays
(39%). The low sensitivity of some of the mutation assays
led to 6 of the 16 being eliminated in Table 3 as opposed
to only 1 of the 7 cytogenetic assays. The normalized (%)
performances of these 2 classes of assays as complementary
tests are compared in Fig. 7. Similar efficiencies are
evident with the chromosomal assays proving marginally more
sensitive to the carcinogens.
Figs. 4, 5, and 7 enable the problem of reproducibility of
response to be visualized and it is clear that harmonization and/or
extension of assay protocols is indicated for both classes of test.
The effort required to produce a common and sensitive assay
protocol, combined with a realistic assessment of whether such
efforts are likely to be rewarded, constitutes the final factor in
the recommendation of the chromosomal-aberration assays. Four
points appear to be particularly relevant:
(a) chromosomal-aberration assays offer an independent genetic
end-point from the Salmonella assay;
(b) chromosomal-aberration assays have a more extensive
history than the mammalian cell gene-mutation assays; this
means that more laboratories have used them for a longer
period; thus, any protocol corrections required should be
more clearly perceived and easier to institute;
(c) the recommendations for assay protocol modification made
by the chromosomal-aberration working group were more
clearly defined, less fundamental, and seemingly easier to
institute than those made by the gene-mutation working
group; and
(d) once the decision is made to conduct a chromosomal-
aberration assay in a laboratory, the expertise and
facilities required will serve equally for the assessment
of aneuploidy, polyploidy, and SCEs.
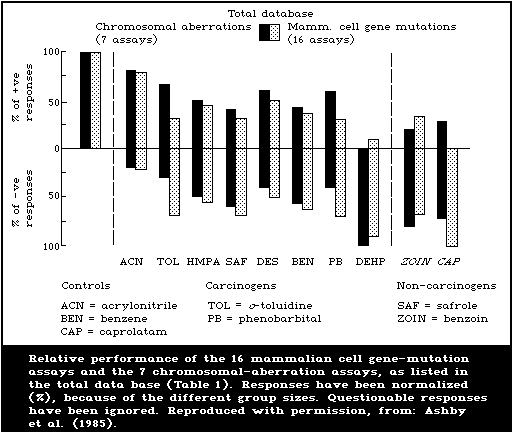 It is therefore recommended that major efforts be focused on
designing a minimum effective protocol according to which
chromosomal-aberration assays should be conducted, and that this
could constitute a complementary assay for the Salmonella test.
Recourse to other assays may be indicated, in order to assess fully
the in vitro activity of certain test chemicals. Acquisition of
test data from a minimum of these 2 standard assays, if optimally
performed, will enable a greater degree of comparability of data
between both independent laboratories and different countries, and
this can only help the efficient detection of possible new human
carcinogens.
In many testing schemes, resources are spread across a large
number of in vitro assays, and this may lead to assays on a new
chemical being conducted suboptimally. The alternative suggested
here is that with 2 well-conducted and common assays, there could
be enhanced comparability and reliability of data, together with an
increase in carcinogen sensitivity. In addition, the Salmonella
assay should be conducted according to what is now regarded as the
best current practice, i.e., employing strains TA1535, 1537, 1538,
98, and 100 in the presence and absence of an induced rat liver S9
mix (10%) (Ames et al., 1975; UKEMS 1983) and also using a
preincubation protocol with strains TA98 and 100 in the case of
initial negative responses. Use of a different source of S9 mix
(e.g., hamster) and additional levels of rat liver S9 mix (e.g., 4%
and 30%) should also be considered (Venitt et al., 1984). In the
case of volatile chemicals, an enclosed test system should be
employed.
The particular cell line employed for cytogenetic assays may be
critical but should not necessarily be restricted to the widely-
used Chinese hamster ovary (CHO) line. Good results were obtained
with the Chinese hamster lung (CHL) line in the CSSTT, but it is
not commercially available at present, and while human lymphocytes
present a sensitive and readily-available source of primary euploid
cells, further studies will be required to demonstrate the general
usefulness of this system (Buckton & Evans, 1982). The Chinese
hamster liver system protocol was designed to measure both
aberrations and aneuploidy and, likewise, will require further
independent studies to confirm the promise shown in the CSSTT.
11. CONCLUSIONS
The dilemma that preceded the formalization of final
conclusions was difficult to resolve. Several carcinogens of
uncertain genotoxicity were selected for the CSSTT, in order to
ascertain which among the many available eukaryotic in vitro
assays for carcinogens would be most efficient and best suited for
adoption as a complement to the generally available Salmonella
mutation assay, by laboratories engaged in the routine screening of
chemicals for potential carcinogens. The organizing committee was
aware of the impact that any conclusions derived from this study
might have on legislative guidelines, but was primarily interested
in the optimal use of the resources of laboratories that may be
attempting to identify possible new human carcinogens. In order to
provide a comprehensive study, the number of types of assays
included was not limited. This non-restrictive method of
collecting assays presented a unique problem; by implication, some
investigators had been invited to submit their assay for scrutiny
for a purpose other than they had originally intended. For
example, many oncologists conduct cell-transformation, aneuploidy,
UDS, etc., assays for research purposes, in order to study the
relationship between genetic end-points and the subtle nuances of
the cancer process. Such research studies may have supported the
selective use of a particular assay that was not claimed to be of
general usefulness for the present purposes, and which was not
being recommended as such by the investigators. It is, therefore,
important to emphasize that the failure of an assay to qualify as a
complementary assay does not preclude its use for research into the
intricate processes of cancer initiation/promotion. This caution
is most important to remember when whole classes of assays such as
"cell transformation", "metabolic cooperation", etc., have been
dismissed from consideration in this collaborative study. On the
one hand, is the immediate and worldwide need for efficient and
reliable means for screening new chemicals, leading to their
registration as safe new products (Ashby et al., 1983); on the
other, the need to use selected in vitro assays to probe the
etiology of chemically-induced cancer. The present study is aimed
solely at the former endeavour, but its findings might expedite the
achievement of the latter. Ancillary findings of the CSSTT are
first, that certain classes of in vitro assays appear to be
insensitive to the cancer-related activities of certain chemical
carcinogens. Second, it is suggested that a clear distinction
should be made between the use of eukaryotic in vitro assays to
confirm genotoxic activity, seen previously for a chemical in
bacteria, and their use to complement bacterial-mutation assays,
i.e., to detect genotoxins found inactive in bacteria; the choice
of assay and the interpretation of data are influenced by the need
originally perceived.
1. Significant differences exist among individual investigators
conducting nominally identical assays. A direct result of the
CSSTT is that many recommended protocol changes were identified
for all assays and some have already been instituted by a
number of investigators. Inadequate test protocols clearly
contributed to the fact that some assays appeared to be
insensitive. It is likely that the need, currently perceived,
to conduct a range of genotoxicity assays on a new chemical has
been stimulated, in part, by the inadequacy of some test
protocols.
2. Urgent attention should be given to the definition of criteria
for judging positive and negative test responses in all assays.
Methods by which weak responses can be interpreted with
confidence were identified, including the adoption of
appropriately sensitive test protocols, the acquisition of a
sufficiently large control data base, and the use of
appropriate statistical methods for data assessment.
3. Carcinogens that are inactive or difficult to detect in the
Salmonella assay fall into 2 distinct groups in this study.
The first include genotoxins that are probably only non-
mutagenic to Salmonella because of protocol deficiencies
associated with the overall metabolic capacity of the assay
system. These agents, HMPA, o-toluidine, safrole, and
acrylonitrile, were detected in most of the eukaryotic assays
studied. Thus, a range of assays can act as limited
complements to the Salmonella mutation assay. The other group
of carcinogens, benzene, DEHP, DES, and phenobarbital,
possessed a more selective range of genotoxic activities, and
none of the assays was selectively sensitive to them. Of the
assays studied, only the induction of chromosomal aberrations,
transformation, gene mutations in mammalian cells, and
aneuploidy in yeast gave encouraging overall performances for
the 8 carcinogens (Conclusion 9).
4. A surprising finding of the CSSTT was the apparent mutagenicity
observed for the 3 carcinogens, benzene, DES, and phenobarbital
in some of the mammalian cell gene-mutation assays. These
activities were not seen by the majority of investigators in
this assay group and were not generally confirmed in either the
Drosophila or fungal gene-mutation assays. If they prove
repeatable, 3 cell-specific gene mutagens will have been
defined. If they are not repeatable, this will emphasize the
need, already perceived by the assay working group, to improve
the protocol and data assessment processes for this class of
assay. The concomitant monitoring of 2 gene loci in a single-
cell line, and the development of appropriate and common data
assessment procedures represent 2 areas for research with
mammalian cell gene-mutation assays. The precise role of these
assays in routine screening programmes requires clarification;
in particular, whether they are to be used to anticipate
possible mammalian carcinogens or possible mammalian mutagens,
or both. The use of negative results in a mammalian cell gene-
mutation assay to negate positive observations made in the
Salmonella assay may be premature.
5. A range of cell-transformation assays was studied, and the
overall performance of several of them was impressive. The
duplicated studies with the Syrian hamster embryo (SHE) assay
and the mouse C3H10T 1/2 assay gave particularly encouraging
results. Nonetheless, these assays are not recommended for
routine screening purposes, at present, because of the
expertise required to carry them out and because of the
problems that remain with clone selection and judgement of when
chemically-induced transformation has occurred.
6. Both of the non-carcinogens showed low incidences of activity
in vitro, several of which were confirmed and most of which
were weak. The clastogenicity of caprolactam for human
lymphocytes and its somatic cell mutagenicity in Drosophila
were particularly clear-cut. The activities observed for these
agents in certain in vitro assays do not, therefore, correlate
with their rodent carcinogenicity, and thus, they represent
false-positive predictions of carcinogenicity. This underlines
the fact that in vitro assays are of use for predicting, rather
than defining carcinogenic activity.
7. Three of the carcinogens, DES, phenobarbital, and DEHP, were
chosen to evaluate the hypothesis that some chemicals can
induce tumours in rodents without first modifying the integrity
of nuclear DNA. Although these chemicals were generally less
active in the tests than the other carcinogens, they each
displayed a range of genotoxic activities in vitro. DEHP
showed least activity, with levels similar to those observed
for the 2 non-carcinogens. Nevertheless, it induced
significant aneuploidy and cell-transformation effects in an
almost total absence of evidence of a direct interaction with
DNA. The data generated for DES confirmed earlier reports that
it is an aneuploidy-inducing and cell-transforming agent, and
the present study also revealed that it was capable of causing
primary damage to DNA; it cannot, therefore, be regarded as
non-genotoxic. Phenobarbital was perhaps the previously best
documented non-genotoxin of those studied, but it has been
defined herein as a gene mutagen, a clastogen, and an
aneuploidy-inducing agent. It is concluded that the phrase
"non-genotoxic" should only be used when a sufficiently large
genotoxicity data base has been collected.
8. Many of the assays evaluated may have an important research
role. However, if their main function is to act as
confirmatory or complementary assays in conjunction with the
Salmonella test, then substantial redundancy among in vitro
eukaryotic genotoxicity assays is evident.
9. In relation to the stated aim of the CSSTT, it is recommended
that resources be applied to the definition of a generally
acceptable and applicable protocol for the conduct of
chromosomal-aberration assays, preferably in an agreed cell
type. Use of this class of assay in conjunction with an
adequate assessment of the mutagenicity of a chemical for
Salmonella should provide an efficient primary screen for
possible new carcinogens. Certain chemicals that induce
tumours in rodents at elevated dose levels will remain
undetected by these tests, as exemplified by DEHP in the
present study. The "detection", in vitro, of such agents may
be achieved by the incorporation of additional assays, but the
price in terms of a reduction in overall specificity and the
increased burden on technical resources may be high.
Consideration of whether the joint inactivity of a chemical in
an adequate Salmonella assay and an adequate in vitro
cytogenetic assay provides evidence of its inactivity with DNA
and, thus, its probable non-carcinogenicity for mammals, at low
dose levels, is indicated. The adoption of a chromosomal-
aberration assay as a common complementary test would have the
additional advantages of allowing easy comparison of data,
ready access to supplementary cytogenetic assays, and the
provision of test data derived using a genetic end-point,
independent of that of the Salmonella (gene-mutation) assay.
REFERENCES
AMES, B.N., MCCANN, J., & YAMASAKI, E. (1975) Methods for
detecting carcinogens and mutagens with the salmonella/mammalian-
microsome mutagenicity test. Mutat. Res., 31: 347-364.
ASHBY, J. (1983) The distinction between genetic and non-genetic
mechanisms in chemical carcinogenesis. In: Zbinden, G., Cohadon,
F., Detaille, J.Y., & Mazué, G., ed. Current problems in drug
toxicity, London, Paris, John Libby Eurotext.
ASHBY, J. & LEFEVRE, P.A. (1983) Genetic toxicology with
formaldehyde and closely related chemicals. In: Gibson, J.E., ed.
Formaldehyde toxicity, Hemisphere Pub.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr, MATTER,
B.E., & SHELBY, M. (1983) The two IPCS collaborative studies on
short-term tests for genotoxicity and carcinogenicity. Mutat. Res.,
109: 123-126.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr,
MARGOLIN, B.H., MATTER, B.E., & SHELBY, M., ed. (1985) Evaluation
of short-term tests for carcinogens. Report of the International
Programme on Chemical Safety Collaborative Study on in vitro
assays, Amsterdam, Elsevier Science Publishers (Progress in
Mutation Research, Vol. 5).
ATHANASIOU, K. & KRYRTOPOULOS, S.A. (1981) Induction of SCE by
non-mutagenic carcinogens. NATO Adv. Study Inst. Ser A, 40: 557-562
(Chem. abst., 97 18839 (1982).
BARRETT, J.C., WONG, A., & MCLACHLAN, J.A. (1981)
Diethylstilboestrol induced neoplastic transformation without
measurable gene mutation at two loci. Science, 212: 1402-1404.
BUCKTON, K.E. & EVANS, H.J. (1982) Human peripheral blood
lymphocyte cultures. An in vitro assay for the cytogenetic effects
of environmental mutagens. In: Hsu, T.C., ed. Cytogenetic assays of
environmental mutagens, Osman, New Jersey, Allanheld.
CAIRNS, J. (1981) The origin of human cancers. Nature (Lond.),
289: 353-357.
CLEMMENSEN, J. & HJALGRIM-JENSEN, S. (1977) On the absence of
carcinogenicity to man of phenobarbital. In: Statistical studies in
malignant neoplasms, Vol. 5, pp. 38-50.
DEAN, B.J. (1978) Genetic toxicology of benzene, toluene,
xylenes, and phenols. Mutat. Res., 47: 75-97.
DE SERRES, F.J. & ASHBY, J., ed. (1981) Evaluation of short-term
tests for carcinogens: report of the international collaborative
program, North-Holland, New York, Elsevier.
GOLDMACHER, V.S. & THILLY, W.G. (1983) Formaldehyde is mutagenic
for cultured human cells. Mutat. Res., 116: 417-422.
HUFF, J.E. (1982) Caprolactam: condensation of the carcinogenesis
bioassay technical report. Environ. Health Perspect., 45: 199-200.
KLEIN, G. (1981) The role of gene dosage and genetic
transpositions in carcinogenesis. Nature (Lond.), 294: 313-318.
MILVY, P. & WOLFF, M. (1977) Mutagenic studies with
acrylonitrile. Mutat. Res., 48: 217-278.
MOODY, D.E. & REDDY, J.K. (1978) Hepatic peroxisome (microbody)
proliferation in rats fed plasticizers and related compounds.
Toxicol. appl. Pharmacol., 45: 497-504.
PHILLIPS, B.J., JAMES, T.E.B., & GANGOLLI, S.D. (1982)
Genotoxicity studies of di(2-ethylhexyl)phthalate and its
metabolities in CHO cells. Mutat. Res., 102: 297-304.
REDDY, E.P., REYNOLDS, R.K., SANTOS, E., & BARBACID, M. (1982) A
point of mutation is responsible for the acquisition of
transforming properties by the T24 human bladder carcinoma
oncogene. Nature (Lond.), 300: 149-152.
STYLES, J.A (1977) A method for detecting carcinogenic organic
chemicals using mammalian cells in culture. Br. J. Cancer, 36: 558-563.
TABIN, C.J., BRADLEY, S.M., BARGMANN, C.I., WEINBERG, R.A.,
PAPAGEORGE, A.G., SCOLNICK, E.M., DHAR, R., LOWY, D.R., & CHANG,
E.H. (1982) Mechanism of activation of a human oncogene. Nature
(Lond.), 300: 143-149.
TOMITA, I., NAKAMURA, Y., AOKI, N., & INUI, N. (1982)
Mutagenic/carcinogenic potential of DEHP and MEHP. Environ. Health
Perspect., 45: 119-125.
UKEMS (1983) Report of the UKEMS sub-committee on guidelines for
mutagenicity testing, Part I, The United Kingdom Environmental
Mutagen Society.
UNITED STATES PUBLIC LAW (1958) United States Code, title 21,
section 348(c)(3)(A).
US NTP (1980) Benzoin - NTP technical report on the toxicology
and carcinogencity of benzoin, Research Triangle Park, North
Carolina, National Toxicology Program (TR No. 204).
US NTP (1982) Caprolactam - NTP technical report on the
toxicology and carcinogenicity of benzoin, Research Triangle Park,
North Carolina, National Toxicology Program (TR No. 214).
VENITT, S., BUSHELL, C.T., & OSBORNE, M. (1977) Mutagenicity of
acrylonitrile (cyanoethylene) in Escherichia coli. Mutat. Res., 45:
283-288.
VENITT, S., CROFTON-SLEIGH, C., & OSBORNE, M.B. (1984) The hair
dye reagent 2-(2,4-diaminophenoxy) ethanol is mutagenic to
Salmonella typhimurium. Mutat. Res., 135: 31-47.
WEINSTEIN, I.B., WIGLER, M., & STADLER, U. (1976) Analysis of the
mechanism of chemical carcinogenesis of epithelial cell cultures,
Lyons, International Agency for Research on Cancer, (IARC
Scientific Publication No. 12, Screening tests in Chemical
Carcinogenesis).
WEISS, R. (1982) The myconcogene in man and birds. Nature
(Lond.), 299: 9-10.
WILLIAMS, G.M. (1981) Liver carcinogenesis: the role for some
chemicals of an epigenetic mechanism of liver-tumour promotion
involving modification of the cell membrane. Food Cosmet. Toxicol.,
19: 577-583.
It is therefore recommended that major efforts be focused on
designing a minimum effective protocol according to which
chromosomal-aberration assays should be conducted, and that this
could constitute a complementary assay for the Salmonella test.
Recourse to other assays may be indicated, in order to assess fully
the in vitro activity of certain test chemicals. Acquisition of
test data from a minimum of these 2 standard assays, if optimally
performed, will enable a greater degree of comparability of data
between both independent laboratories and different countries, and
this can only help the efficient detection of possible new human
carcinogens.
In many testing schemes, resources are spread across a large
number of in vitro assays, and this may lead to assays on a new
chemical being conducted suboptimally. The alternative suggested
here is that with 2 well-conducted and common assays, there could
be enhanced comparability and reliability of data, together with an
increase in carcinogen sensitivity. In addition, the Salmonella
assay should be conducted according to what is now regarded as the
best current practice, i.e., employing strains TA1535, 1537, 1538,
98, and 100 in the presence and absence of an induced rat liver S9
mix (10%) (Ames et al., 1975; UKEMS 1983) and also using a
preincubation protocol with strains TA98 and 100 in the case of
initial negative responses. Use of a different source of S9 mix
(e.g., hamster) and additional levels of rat liver S9 mix (e.g., 4%
and 30%) should also be considered (Venitt et al., 1984). In the
case of volatile chemicals, an enclosed test system should be
employed.
The particular cell line employed for cytogenetic assays may be
critical but should not necessarily be restricted to the widely-
used Chinese hamster ovary (CHO) line. Good results were obtained
with the Chinese hamster lung (CHL) line in the CSSTT, but it is
not commercially available at present, and while human lymphocytes
present a sensitive and readily-available source of primary euploid
cells, further studies will be required to demonstrate the general
usefulness of this system (Buckton & Evans, 1982). The Chinese
hamster liver system protocol was designed to measure both
aberrations and aneuploidy and, likewise, will require further
independent studies to confirm the promise shown in the CSSTT.
11. CONCLUSIONS
The dilemma that preceded the formalization of final
conclusions was difficult to resolve. Several carcinogens of
uncertain genotoxicity were selected for the CSSTT, in order to
ascertain which among the many available eukaryotic in vitro
assays for carcinogens would be most efficient and best suited for
adoption as a complement to the generally available Salmonella
mutation assay, by laboratories engaged in the routine screening of
chemicals for potential carcinogens. The organizing committee was
aware of the impact that any conclusions derived from this study
might have on legislative guidelines, but was primarily interested
in the optimal use of the resources of laboratories that may be
attempting to identify possible new human carcinogens. In order to
provide a comprehensive study, the number of types of assays
included was not limited. This non-restrictive method of
collecting assays presented a unique problem; by implication, some
investigators had been invited to submit their assay for scrutiny
for a purpose other than they had originally intended. For
example, many oncologists conduct cell-transformation, aneuploidy,
UDS, etc., assays for research purposes, in order to study the
relationship between genetic end-points and the subtle nuances of
the cancer process. Such research studies may have supported the
selective use of a particular assay that was not claimed to be of
general usefulness for the present purposes, and which was not
being recommended as such by the investigators. It is, therefore,
important to emphasize that the failure of an assay to qualify as a
complementary assay does not preclude its use for research into the
intricate processes of cancer initiation/promotion. This caution
is most important to remember when whole classes of assays such as
"cell transformation", "metabolic cooperation", etc., have been
dismissed from consideration in this collaborative study. On the
one hand, is the immediate and worldwide need for efficient and
reliable means for screening new chemicals, leading to their
registration as safe new products (Ashby et al., 1983); on the
other, the need to use selected in vitro assays to probe the
etiology of chemically-induced cancer. The present study is aimed
solely at the former endeavour, but its findings might expedite the
achievement of the latter. Ancillary findings of the CSSTT are
first, that certain classes of in vitro assays appear to be
insensitive to the cancer-related activities of certain chemical
carcinogens. Second, it is suggested that a clear distinction
should be made between the use of eukaryotic in vitro assays to
confirm genotoxic activity, seen previously for a chemical in
bacteria, and their use to complement bacterial-mutation assays,
i.e., to detect genotoxins found inactive in bacteria; the choice
of assay and the interpretation of data are influenced by the need
originally perceived.
1. Significant differences exist among individual investigators
conducting nominally identical assays. A direct result of the
CSSTT is that many recommended protocol changes were identified
for all assays and some have already been instituted by a
number of investigators. Inadequate test protocols clearly
contributed to the fact that some assays appeared to be
insensitive. It is likely that the need, currently perceived,
to conduct a range of genotoxicity assays on a new chemical has
been stimulated, in part, by the inadequacy of some test
protocols.
2. Urgent attention should be given to the definition of criteria
for judging positive and negative test responses in all assays.
Methods by which weak responses can be interpreted with
confidence were identified, including the adoption of
appropriately sensitive test protocols, the acquisition of a
sufficiently large control data base, and the use of
appropriate statistical methods for data assessment.
3. Carcinogens that are inactive or difficult to detect in the
Salmonella assay fall into 2 distinct groups in this study.
The first include genotoxins that are probably only non-
mutagenic to Salmonella because of protocol deficiencies
associated with the overall metabolic capacity of the assay
system. These agents, HMPA, o-toluidine, safrole, and
acrylonitrile, were detected in most of the eukaryotic assays
studied. Thus, a range of assays can act as limited
complements to the Salmonella mutation assay. The other group
of carcinogens, benzene, DEHP, DES, and phenobarbital,
possessed a more selective range of genotoxic activities, and
none of the assays was selectively sensitive to them. Of the
assays studied, only the induction of chromosomal aberrations,
transformation, gene mutations in mammalian cells, and
aneuploidy in yeast gave encouraging overall performances for
the 8 carcinogens (Conclusion 9).
4. A surprising finding of the CSSTT was the apparent mutagenicity
observed for the 3 carcinogens, benzene, DES, and phenobarbital
in some of the mammalian cell gene-mutation assays. These
activities were not seen by the majority of investigators in
this assay group and were not generally confirmed in either the
Drosophila or fungal gene-mutation assays. If they prove
repeatable, 3 cell-specific gene mutagens will have been
defined. If they are not repeatable, this will emphasize the
need, already perceived by the assay working group, to improve
the protocol and data assessment processes for this class of
assay. The concomitant monitoring of 2 gene loci in a single-
cell line, and the development of appropriate and common data
assessment procedures represent 2 areas for research with
mammalian cell gene-mutation assays. The precise role of these
assays in routine screening programmes requires clarification;
in particular, whether they are to be used to anticipate
possible mammalian carcinogens or possible mammalian mutagens,
or both. The use of negative results in a mammalian cell gene-
mutation assay to negate positive observations made in the
Salmonella assay may be premature.
5. A range of cell-transformation assays was studied, and the
overall performance of several of them was impressive. The
duplicated studies with the Syrian hamster embryo (SHE) assay
and the mouse C3H10T 1/2 assay gave particularly encouraging
results. Nonetheless, these assays are not recommended for
routine screening purposes, at present, because of the
expertise required to carry them out and because of the
problems that remain with clone selection and judgement of when
chemically-induced transformation has occurred.
6. Both of the non-carcinogens showed low incidences of activity
in vitro, several of which were confirmed and most of which
were weak. The clastogenicity of caprolactam for human
lymphocytes and its somatic cell mutagenicity in Drosophila
were particularly clear-cut. The activities observed for these
agents in certain in vitro assays do not, therefore, correlate
with their rodent carcinogenicity, and thus, they represent
false-positive predictions of carcinogenicity. This underlines
the fact that in vitro assays are of use for predicting, rather
than defining carcinogenic activity.
7. Three of the carcinogens, DES, phenobarbital, and DEHP, were
chosen to evaluate the hypothesis that some chemicals can
induce tumours in rodents without first modifying the integrity
of nuclear DNA. Although these chemicals were generally less
active in the tests than the other carcinogens, they each
displayed a range of genotoxic activities in vitro. DEHP
showed least activity, with levels similar to those observed
for the 2 non-carcinogens. Nevertheless, it induced
significant aneuploidy and cell-transformation effects in an
almost total absence of evidence of a direct interaction with
DNA. The data generated for DES confirmed earlier reports that
it is an aneuploidy-inducing and cell-transforming agent, and
the present study also revealed that it was capable of causing
primary damage to DNA; it cannot, therefore, be regarded as
non-genotoxic. Phenobarbital was perhaps the previously best
documented non-genotoxin of those studied, but it has been
defined herein as a gene mutagen, a clastogen, and an
aneuploidy-inducing agent. It is concluded that the phrase
"non-genotoxic" should only be used when a sufficiently large
genotoxicity data base has been collected.
8. Many of the assays evaluated may have an important research
role. However, if their main function is to act as
confirmatory or complementary assays in conjunction with the
Salmonella test, then substantial redundancy among in vitro
eukaryotic genotoxicity assays is evident.
9. In relation to the stated aim of the CSSTT, it is recommended
that resources be applied to the definition of a generally
acceptable and applicable protocol for the conduct of
chromosomal-aberration assays, preferably in an agreed cell
type. Use of this class of assay in conjunction with an
adequate assessment of the mutagenicity of a chemical for
Salmonella should provide an efficient primary screen for
possible new carcinogens. Certain chemicals that induce
tumours in rodents at elevated dose levels will remain
undetected by these tests, as exemplified by DEHP in the
present study. The "detection", in vitro, of such agents may
be achieved by the incorporation of additional assays, but the
price in terms of a reduction in overall specificity and the
increased burden on technical resources may be high.
Consideration of whether the joint inactivity of a chemical in
an adequate Salmonella assay and an adequate in vitro
cytogenetic assay provides evidence of its inactivity with DNA
and, thus, its probable non-carcinogenicity for mammals, at low
dose levels, is indicated. The adoption of a chromosomal-
aberration assay as a common complementary test would have the
additional advantages of allowing easy comparison of data,
ready access to supplementary cytogenetic assays, and the
provision of test data derived using a genetic end-point,
independent of that of the Salmonella (gene-mutation) assay.
REFERENCES
AMES, B.N., MCCANN, J., & YAMASAKI, E. (1975) Methods for
detecting carcinogens and mutagens with the salmonella/mammalian-
microsome mutagenicity test. Mutat. Res., 31: 347-364.
ASHBY, J. (1983) The distinction between genetic and non-genetic
mechanisms in chemical carcinogenesis. In: Zbinden, G., Cohadon,
F., Detaille, J.Y., & Mazué, G., ed. Current problems in drug
toxicity, London, Paris, John Libby Eurotext.
ASHBY, J. & LEFEVRE, P.A. (1983) Genetic toxicology with
formaldehyde and closely related chemicals. In: Gibson, J.E., ed.
Formaldehyde toxicity, Hemisphere Pub.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr, MATTER,
B.E., & SHELBY, M. (1983) The two IPCS collaborative studies on
short-term tests for genotoxicity and carcinogenicity. Mutat. Res.,
109: 123-126.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr,
MARGOLIN, B.H., MATTER, B.E., & SHELBY, M., ed. (1985) Evaluation
of short-term tests for carcinogens. Report of the International
Programme on Chemical Safety Collaborative Study on in vitro
assays, Amsterdam, Elsevier Science Publishers (Progress in
Mutation Research, Vol. 5).
ATHANASIOU, K. & KRYRTOPOULOS, S.A. (1981) Induction of SCE by
non-mutagenic carcinogens. NATO Adv. Study Inst. Ser A, 40: 557-562
(Chem. abst., 97 18839 (1982).
BARRETT, J.C., WONG, A., & MCLACHLAN, J.A. (1981)
Diethylstilboestrol induced neoplastic transformation without
measurable gene mutation at two loci. Science, 212: 1402-1404.
BUCKTON, K.E. & EVANS, H.J. (1982) Human peripheral blood
lymphocyte cultures. An in vitro assay for the cytogenetic effects
of environmental mutagens. In: Hsu, T.C., ed. Cytogenetic assays of
environmental mutagens, Osman, New Jersey, Allanheld.
CAIRNS, J. (1981) The origin of human cancers. Nature (Lond.),
289: 353-357.
CLEMMENSEN, J. & HJALGRIM-JENSEN, S. (1977) On the absence of
carcinogenicity to man of phenobarbital. In: Statistical studies in
malignant neoplasms, Vol. 5, pp. 38-50.
DEAN, B.J. (1978) Genetic toxicology of benzene, toluene,
xylenes, and phenols. Mutat. Res., 47: 75-97.
DE SERRES, F.J. & ASHBY, J., ed. (1981) Evaluation of short-term
tests for carcinogens: report of the international collaborative
program, North-Holland, New York, Elsevier.
GOLDMACHER, V.S. & THILLY, W.G. (1983) Formaldehyde is mutagenic
for cultured human cells. Mutat. Res., 116: 417-422.
HUFF, J.E. (1982) Caprolactam: condensation of the carcinogenesis
bioassay technical report. Environ. Health Perspect., 45: 199-200.
KLEIN, G. (1981) The role of gene dosage and genetic
transpositions in carcinogenesis. Nature (Lond.), 294: 313-318.
MILVY, P. & WOLFF, M. (1977) Mutagenic studies with
acrylonitrile. Mutat. Res., 48: 217-278.
MOODY, D.E. & REDDY, J.K. (1978) Hepatic peroxisome (microbody)
proliferation in rats fed plasticizers and related compounds.
Toxicol. appl. Pharmacol., 45: 497-504.
PHILLIPS, B.J., JAMES, T.E.B., & GANGOLLI, S.D. (1982)
Genotoxicity studies of di(2-ethylhexyl)phthalate and its
metabolities in CHO cells. Mutat. Res., 102: 297-304.
REDDY, E.P., REYNOLDS, R.K., SANTOS, E., & BARBACID, M. (1982) A
point of mutation is responsible for the acquisition of
transforming properties by the T24 human bladder carcinoma
oncogene. Nature (Lond.), 300: 149-152.
STYLES, J.A (1977) A method for detecting carcinogenic organic
chemicals using mammalian cells in culture. Br. J. Cancer, 36: 558-563.
TABIN, C.J., BRADLEY, S.M., BARGMANN, C.I., WEINBERG, R.A.,
PAPAGEORGE, A.G., SCOLNICK, E.M., DHAR, R., LOWY, D.R., & CHANG,
E.H. (1982) Mechanism of activation of a human oncogene. Nature
(Lond.), 300: 143-149.
TOMITA, I., NAKAMURA, Y., AOKI, N., & INUI, N. (1982)
Mutagenic/carcinogenic potential of DEHP and MEHP. Environ. Health
Perspect., 45: 119-125.
UKEMS (1983) Report of the UKEMS sub-committee on guidelines for
mutagenicity testing, Part I, The United Kingdom Environmental
Mutagen Society.
UNITED STATES PUBLIC LAW (1958) United States Code, title 21,
section 348(c)(3)(A).
US NTP (1980) Benzoin - NTP technical report on the toxicology
and carcinogencity of benzoin, Research Triangle Park, North
Carolina, National Toxicology Program (TR No. 204).
US NTP (1982) Caprolactam - NTP technical report on the
toxicology and carcinogenicity of benzoin, Research Triangle Park,
North Carolina, National Toxicology Program (TR No. 214).
VENITT, S., BUSHELL, C.T., & OSBORNE, M. (1977) Mutagenicity of
acrylonitrile (cyanoethylene) in Escherichia coli. Mutat. Res., 45:
283-288.
VENITT, S., CROFTON-SLEIGH, C., & OSBORNE, M.B. (1984) The hair
dye reagent 2-(2,4-diaminophenoxy) ethanol is mutagenic to
Salmonella typhimurium. Mutat. Res., 135: 31-47.
WEINSTEIN, I.B., WIGLER, M., & STADLER, U. (1976) Analysis of the
mechanism of chemical carcinogenesis of epithelial cell cultures,
Lyons, International Agency for Research on Cancer, (IARC
Scientific Publication No. 12, Screening tests in Chemical
Carcinogenesis).
WEISS, R. (1982) The myconcogene in man and birds. Nature
(Lond.), 299: 9-10.
WILLIAMS, G.M. (1981) Liver carcinogenesis: the role for some
chemicals of an epigenetic mechanism of liver-tumour promotion
involving modification of the cell membrane. Food Cosmet. Toxicol.,
19: 577-583.
 The performance of assays on the reduced list was carefully
assessed by the Editorial Committee, as it was felt that, for the
guidance of the reader, some overall evaluation of this complex
project was necessary, even though at this stage the opinions were
those of scientists, closely associated with the project, and,
thus, might be subjective. Close attention was paid to the
conclusions drawn by the assay groups regarding the usefulness of
individual assays. However, not all of these working groups
approached an assessment of their assays within the context
outlined originally. On occasion, the recommendations and
conclusions derived below are at variance with conclusions of the
group but, generally, they are consistent with them or represent
the only ones available.
9.1. Gene Mutation in Yeast
These assays are, phylogenically, the nearest to the Salmonella
assay and share a similar genetic end-point. Previous studies have
also established that genotoxic agents, found active in Salmonella,
are generally also active in the corresponding yeast gene-mutation
assays. It was, therefore, expected that these 2 classes of assay
would share similar sensitivities. Fourteen positive responses
were observed within the group comprising HMPA, o-toluidine,
safrole, and acrylonitrile, while only 3 positive responses were
observed for benzene, DEHP, DES, and phenobarbital, and there was
general agreement on the gene mutagenicity of HMPA (4/5 positive)
(Table 3, Fig. 2). The enhanced sensitivity of the yeast gene-
mutation assays, as opposed to the Salmonella mutation assay, to
the 4 assumed genotoxins may be due to endogenous metabolism,
especially evident when the cells are in active growth. It was,
therefore, concluded that this class of assay might represent a
possible alternative to the Salmonella assay, and might present
advantages, on occasion, because of its enhanced sensitivity to
certain genotoxins. The mitochondrial mutation assay (2.1.9) has
been included in this discussion, but should correctly be regarded
as a distinct assay. Two potential problems are presented by these
assays, and by yeast assays in general. First, they are not yet
conducted according to an agreed protocol. For example, the data
shown in Table 2 represent the consensus gained after conducting
assays within certain laboratories under a range of test
conditions: low or high glucose, stationary, semi-stationary, and
log-phase cells, etc. In addition, the largely positive data base
of assay 2.1.4 includes 2 classes of positive response (see assay
group report and investigator report). Second, the Salmonella
assay must remain the preferred primary assay for detecting gene
mutagens in vitro, owing to the extensive data base available and
the large number of laboratories involved (ca 2000).
It was concluded that appropriate yeast assays present a useful
method of detecting gene mutagens, and, on occasion, can be more
sensitive than the Salmonella test. However, given the current
state of the art, they would not be recommended as primary gene-
mutation assays and their usefulness as a single complement to the
Salmonella test is reduced by the repetition of the genetic end-
point involved. The preferential detection by these assays of
HMPA, o-toluidine, safrole, and acrylonitrile is referred to later.
9.2. Drosophila Somatic Cell Mutation Assays
New somatic cell mutation assays have recently been developed
in Drosophila. Their novelty is evidenced by the fact that data
supporting their general sensitivity to genotoxic agents (i.e.,
agents readily detected by the Salmonella assay) are only now
being collected for publication. General sensitivity is, however,
claimed (see working group report). Two different types of assay
were represented. Assays 3.1.1 and 3.1.3, although based in
different tissues, were capable of detecting a range of induced
genetic events, i.e., chromosomal breakage, deletions and
aneuploidy, gene mutations, and mitotic recombinations. These 2
assays were, in fact, more sensitive to the present test chemicals
than assay 3.1.2, where an inappropriate route of administration
was employed and the range of detected mutagenic events was limited
to gene mutations and deletions. Nevertheless, the general
concordance between the responses recorded in each of these 3
assays suggests that they are likely to yield reproducible data
between laboratories.
The performance of assays on the reduced list was carefully
assessed by the Editorial Committee, as it was felt that, for the
guidance of the reader, some overall evaluation of this complex
project was necessary, even though at this stage the opinions were
those of scientists, closely associated with the project, and,
thus, might be subjective. Close attention was paid to the
conclusions drawn by the assay groups regarding the usefulness of
individual assays. However, not all of these working groups
approached an assessment of their assays within the context
outlined originally. On occasion, the recommendations and
conclusions derived below are at variance with conclusions of the
group but, generally, they are consistent with them or represent
the only ones available.
9.1. Gene Mutation in Yeast
These assays are, phylogenically, the nearest to the Salmonella
assay and share a similar genetic end-point. Previous studies have
also established that genotoxic agents, found active in Salmonella,
are generally also active in the corresponding yeast gene-mutation
assays. It was, therefore, expected that these 2 classes of assay
would share similar sensitivities. Fourteen positive responses
were observed within the group comprising HMPA, o-toluidine,
safrole, and acrylonitrile, while only 3 positive responses were
observed for benzene, DEHP, DES, and phenobarbital, and there was
general agreement on the gene mutagenicity of HMPA (4/5 positive)
(Table 3, Fig. 2). The enhanced sensitivity of the yeast gene-
mutation assays, as opposed to the Salmonella mutation assay, to
the 4 assumed genotoxins may be due to endogenous metabolism,
especially evident when the cells are in active growth. It was,
therefore, concluded that this class of assay might represent a
possible alternative to the Salmonella assay, and might present
advantages, on occasion, because of its enhanced sensitivity to
certain genotoxins. The mitochondrial mutation assay (2.1.9) has
been included in this discussion, but should correctly be regarded
as a distinct assay. Two potential problems are presented by these
assays, and by yeast assays in general. First, they are not yet
conducted according to an agreed protocol. For example, the data
shown in Table 2 represent the consensus gained after conducting
assays within certain laboratories under a range of test
conditions: low or high glucose, stationary, semi-stationary, and
log-phase cells, etc. In addition, the largely positive data base
of assay 2.1.4 includes 2 classes of positive response (see assay
group report and investigator report). Second, the Salmonella
assay must remain the preferred primary assay for detecting gene
mutagens in vitro, owing to the extensive data base available and
the large number of laboratories involved (ca 2000).
It was concluded that appropriate yeast assays present a useful
method of detecting gene mutagens, and, on occasion, can be more
sensitive than the Salmonella test. However, given the current
state of the art, they would not be recommended as primary gene-
mutation assays and their usefulness as a single complement to the
Salmonella test is reduced by the repetition of the genetic end-
point involved. The preferential detection by these assays of
HMPA, o-toluidine, safrole, and acrylonitrile is referred to later.
9.2. Drosophila Somatic Cell Mutation Assays
New somatic cell mutation assays have recently been developed
in Drosophila. Their novelty is evidenced by the fact that data
supporting their general sensitivity to genotoxic agents (i.e.,
agents readily detected by the Salmonella assay) are only now
being collected for publication. General sensitivity is, however,
claimed (see working group report). Two different types of assay
were represented. Assays 3.1.1 and 3.1.3, although based in
different tissues, were capable of detecting a range of induced
genetic events, i.e., chromosomal breakage, deletions and
aneuploidy, gene mutations, and mitotic recombinations. These 2
assays were, in fact, more sensitive to the present test chemicals
than assay 3.1.2, where an inappropriate route of administration
was employed and the range of detected mutagenic events was limited
to gene mutations and deletions. Nevertheless, the general
concordance between the responses recorded in each of these 3
assays suggests that they are likely to yield reproducible data
between laboratories.
 Assays using Drosophila require minimal technical facilities
but may prove relatively time-consuming. They present the appeal
of conducting an assay on a multicellular organism with its
inherent metabolic capability. It cannot be automatically assumed
that this capability is significantly related to that of a mammal,
but the sensitivity of these assays to a range of genotoxic agents
is consistent with their possession of a broad range of metabolic
pathways.
The 2 most sensitive assays (3.1.1 and 3.1.3) detected 5 and
perhaps 6 of the test agents, one of which was the non-carcinogen
caprolactam (see subsequent discussion of CAP and ZOIN). Activity
was mainly centred in the first group of carcinogens (Table 4, Fig.
3). This performance suggests that somatic mutation in Drosophila,
especially the assays capable of detecting a wide range of genetic
events, presents a promising new assay. However, their
preferential detection of HMPA, o-toluidine, safrole, and
acrylonitrile reduces their value as a general complementary assay
(cf. similar discussion of yeast gene-mutation assays, SCE assays,
and final conclusions). The present discussion is only related to
the use of these assays for predicting the possible mammalian
carcinogenicity of an agent; their use for predicting in vivo
mammalian mutagenic events should be considered separately (see
also discussions of mammalian cell gene-mutation assays).
Assays using Drosophila require minimal technical facilities
but may prove relatively time-consuming. They present the appeal
of conducting an assay on a multicellular organism with its
inherent metabolic capability. It cannot be automatically assumed
that this capability is significantly related to that of a mammal,
but the sensitivity of these assays to a range of genotoxic agents
is consistent with their possession of a broad range of metabolic
pathways.
The 2 most sensitive assays (3.1.1 and 3.1.3) detected 5 and
perhaps 6 of the test agents, one of which was the non-carcinogen
caprolactam (see subsequent discussion of CAP and ZOIN). Activity
was mainly centred in the first group of carcinogens (Table 4, Fig.
3). This performance suggests that somatic mutation in Drosophila,
especially the assays capable of detecting a wide range of genetic
events, presents a promising new assay. However, their
preferential detection of HMPA, o-toluidine, safrole, and
acrylonitrile reduces their value as a general complementary assay
(cf. similar discussion of yeast gene-mutation assays, SCE assays,
and final conclusions). The present discussion is only related to
the use of these assays for predicting the possible mammalian
carcinogenicity of an agent; their use for predicting in vivo
mammalian mutagenic events should be considered separately (see
also discussions of mammalian cell gene-mutation assays).
 9.3. Assays for DNA Damage SSB (Single-Strand Breaks) and UDS
(Detected via Autoradiography or Scintillation Counting)
Although both of these classes of assay are designed to be
sensitive to the consequences of the primary interaction of the
test chemical (or a metabolite) with DNA, there are good reasons
for not considering them as equivalent. These reasons are
discussed in the assay working group report and are supported by
aspects of the present data base (Tables 2 and 3). For example,
HMPA was inactive in the 3 SSB assays yet was active in the assays
for UDS (2+ and 1?). In contrast, DES was uniformly active in the
3 SSB assays and inactive in the UDS tests. The fact that
unanimity between these 2 assay groups can be achieved is evidenced
by the responses seen for TOL (6/6+), SAF (5/6+), and PB (4/4-).
The relative sensitivity of these 3 classes of assay are shown in
Fig. 3. It should be noted that some legislative authorities have
discussed the need for assay data indicating primary interaction of
a test chemical with DNA. Their requirement could be met with data
generated from either of these groups of assay, i.e., the 2 are
generally regarded as being different methods of assaying a similar
phenomenon. As can be seen, this may not be so.
Perhaps one of the most surprising findings of the CSSTT is the
insensitivity demonstrated by the 2 autoradiographic assays (4.4.1
and 4.4.2, eliminated from Table 4). These were conducted
according to similar protocols and are well represented in earlier
literature. Their poor performance in the present study suggests
that they are unsuitable for use as a complementary assay, though
their use for confirming the activity in mammalian cells of
previously defined bacterial mutagens remains. Another intriguing
aspect of their poor performance is that it provides evidence to
suggest that the autoradiographic end-point is less sensitive than
the other end-points under review (with the exception of the
activity seen for HMPA in one assay, 4.4.1). This insensitivity is
unlikely to be due to metabolic incompetence as the hepatocytes
employed were primary isolates, the metabolic competence of which
had been previously established. Furthermore, the possibility that
this divergence of data was due to these agents producing only
short-patch repair is reduced, given the range of chemically
different carcinogens employed. Whether some of the positive
responses observed in the SSB assays were a direct result of cell
necrosis is discussed below.
Assays for SSB and UDS would be expected to be primarily
sensitive to the 4 confirmed genotoxins represented here, HMPA,
o-toluidine, safrole, and acrylonitrile, and this was observed (cf.
Table 3 and the discussion of the yeast and Drosophila gene-
mutation assays in this section). The only confirmed success with
the remaining 4 carcinogens was the ability of all of the SSB
assays to detect DES. Non-reproducible activity was seen in both
classes of assay for benzene (2/6), benzoin (2/5), and DEHP (1/5).
The latter isolated positive response was strongly endorsed by the
investigator (4.4.3) and requires further investigation.
A topic for future research is the extent to which cell
necrosis, with the consequent formation of broken DNA, contributes
to the outcome of SSB assays. For example, the isolated positive
response observed for benzene (assay 4.3.2) was only observed at
higher dose levels than those employed by the other 2 investigators
in this group. This response may, therefore, represent a technical
false-positive response induced solely by DNA strand breakage as
the result of toxicity. Equally, the extent to which
"mitotic" DNA synthesis has been successfully blocked during the
conduct of a scintillation UDS assay relates directly to the
credibility accorded to weak induced effects. The above 2 concerns
may explain some of the divergencies of data evident within these 3
classes of assay for the present chemicals.
Given the simple requirement to provide evidence of the ability
of a chemical to induce primary DNA damage in mammalian cells, the
investigator is faced with the following decisions:
(a) If the agent is already defined as a bacterial mutagen,
any of the 3 classes of assay (SSB or UDS via
autoradiography or scintillation counting) discussed
herein could be used.
(b) If the agent is inactive as a bacterial mutagen, use of
either the SSB or scintillation counted UDS assays could
present advantages, depending on the chemicals involved.
The results obtained for HMPA and DES suggest that the use
of both assays should be considered, if an assay for "DNA
damage" is required as a complement to bacterial
mutagenicity.
It was, therefore, concluded that the simple legislative
requirement for "evidence of DNA damage" was too vague to be
meaningful. If complementary data are being sought, the
autoradiographic end-point should be avoided and data should be
derived from an SSB assay and/or scintillation counted UDS assay.
The failure of the CSSTT to indicate clearly a single class of
assay, within this group, for routine adoption suggests that
attention should be given to developing the sensitivity of DNA
damage assays to the point that only a single test is required.
The relative performance of these 3 classes of DNA repair assays is
shown in Fig. 3.
9.4. Assays for the Induction of Aneuploidy
In the past, assays for the induction of aneuploidy have not
generally been considered as predictive tests for carcinogenicity.
In the first place, few appropriate in vitro assays existed, and
secondly, the induction of aneuploidy was assumed to have greater
relevance for the promotional, as opposed to the initiatory stages
of carcinogenesis. Thus, while the previous data base has included
the evaluation of a few reference genotoxins such as MNU, it has
mainly focused on cancer-promoting agents such as the phorbol
esters and saccharin, etc. Two investigators have provided the
greater part of the present data base for this class of assay by
presenting data derived from yeast assays (2.4.1.3). In addition,
data derived using mammalian cells were presented (4.9.1) together
with a set of data observed in Aspergillus (2.4.4).
The performance of the aneuploidy assays in the CSSTT was
impressive (Table 2 and 3). In the case of the 2 sets of yeast
data provided from one laboratory (2.4.1 and 2.4.3), a 100%
correlation between the reported carcinogenicity of the test agents
and their activity in vitro was observed. In particular, the
unanimous detection of both DEHP and phenobarbital by these
aneuploidy assays is unique in this study.
The primary conclusion regarding this group of assays is,
therefore, that they present high promise for development as a
class of complementary assays. The development phase should be
influenced by the following considerations, some of which are of
sufficient importance to ensure that they are resolved before any
particular aneuploidy assay is recommended for general adoption.
(a) The findings of the CSSTT suggest that the induction of
aneuploidy may be a critical step in the mechanism of
action of complete carcinogens. These observations
require independent confirmation in other laboratories,
for both the yeast and the mammalian cell systems.
(b) In the case of the D61-M yeast data from the 2
laboratories considered, major discrepancies were evident.
These were all of the same nature and represented the
failure of one laboratory to detect anything beyond
benzene and DES. It is possible that this divergence was
due to technical factors affecting the mitotic or
metabolic activity of the yeast. For example, evidence
from the fungal studies supported an earlier observation
that growing cells are more sensitive to genotoxins than
stationary cells. This could be because of the reduced
metabolic activity of stationary cells. Thus, this
divergence might be due to a relatively minor technical
consideration, but it must be defined, understood,
corrected, and defended in the literature before any
general adoption of yeast aneuploidy assays can be
recommended. The question of whether the D6 strain can be
regarded as repetitive of the D61-M strain also requires
resolution; if it is to be retained, its efficacy must be
confirmed in other laboratories.
(c) The isolated set of Aspergillus data suggests that this
assay is relatively insensitive. Clearly, if the
induction of aneuploidy is to be pursued as a useful
complementary end-point, and if Aspergillus nidulans is to
be selected as the marker organism, a separate data base
will have to be generated to justify this choice.
(d) Requirements similar to those in (c) must apply to the
isolated set of aneuploidy data generated in the Chinese
hamster liver fibroblast assay (4.9.1). This assay is
relatively novel; thus, apart from an expansion of the
data base and the commissioning of independent studies,
the following point requires attention. The cells used in
this study were of between the 9 - 15th passage. The
corollary for using such a spread of cell passages is that
sufficient evidence must be presented on the chromosomal
number and distribution in each cell line before test data
can be interpreted with confidence. Also, the multiple
scoring of hyper, hypo, and polyploid cells presents a
complex data base, the statistical interpretation of which
might not yet have been optimized. Furthermore, the
possible karyotypic instability of higher passage cells
suggests that the current practice of scoring only 200
cells each in test and control cultures might not be
adequate.
The aneuploidy findings of the CSSTT, therefore, provide an
exciting area for future research. However, the role of such
assays as routinely employed complementary tests must await the
acquisition of further data and the outcome of developmental
studies. The related polyploidy assays are considered in section
9.7.
9.5. Mammalian Cell Gene-Mutation Assays
Mammalian cell gene-mutation assays, together with chromosomal-
aberration assays, are high on the list of optional secondary
assays referred to by legislative authorities. The performance of
these assays was, thus, of particular interest. The mammalian
gene-mutation assays constituted the largest group of assays in the
project.
Sixteen sets of data were considered, 6 of which were based on
nominally the same assay (L5178Y), employing the same selective
agent (trifluorothymidine).
Qualitative inspection of the total data base for the mammalian
gene-mutation assays (Table 2), and of that produced following the
primary reduction step (Table 4), was not, as pointed out by the
working group, initially encouraging. The overall sensitivity of
the collected gene-mutation and chromosomal-aberration assays were
similar (Table 3; 62 and 67% and 46 and 53% overall sensitivity,
respectively, for the 2 groups of 4 carcinogens). The value of
such numerical indices can be questioned, but they are supported by
the individual comparisons drawn in Table 3 for the activity of
each chemical in these 2 classes of assay. Thus, similar overall
sensitivity was observed in both classes of assay for most of the
chemicals. Fig. 4 and 5 show the comparative sensitivity
histograms with the Salmonella assays.
9.3. Assays for DNA Damage SSB (Single-Strand Breaks) and UDS
(Detected via Autoradiography or Scintillation Counting)
Although both of these classes of assay are designed to be
sensitive to the consequences of the primary interaction of the
test chemical (or a metabolite) with DNA, there are good reasons
for not considering them as equivalent. These reasons are
discussed in the assay working group report and are supported by
aspects of the present data base (Tables 2 and 3). For example,
HMPA was inactive in the 3 SSB assays yet was active in the assays
for UDS (2+ and 1?). In contrast, DES was uniformly active in the
3 SSB assays and inactive in the UDS tests. The fact that
unanimity between these 2 assay groups can be achieved is evidenced
by the responses seen for TOL (6/6+), SAF (5/6+), and PB (4/4-).
The relative sensitivity of these 3 classes of assay are shown in
Fig. 3. It should be noted that some legislative authorities have
discussed the need for assay data indicating primary interaction of
a test chemical with DNA. Their requirement could be met with data
generated from either of these groups of assay, i.e., the 2 are
generally regarded as being different methods of assaying a similar
phenomenon. As can be seen, this may not be so.
Perhaps one of the most surprising findings of the CSSTT is the
insensitivity demonstrated by the 2 autoradiographic assays (4.4.1
and 4.4.2, eliminated from Table 4). These were conducted
according to similar protocols and are well represented in earlier
literature. Their poor performance in the present study suggests
that they are unsuitable for use as a complementary assay, though
their use for confirming the activity in mammalian cells of
previously defined bacterial mutagens remains. Another intriguing
aspect of their poor performance is that it provides evidence to
suggest that the autoradiographic end-point is less sensitive than
the other end-points under review (with the exception of the
activity seen for HMPA in one assay, 4.4.1). This insensitivity is
unlikely to be due to metabolic incompetence as the hepatocytes
employed were primary isolates, the metabolic competence of which
had been previously established. Furthermore, the possibility that
this divergence of data was due to these agents producing only
short-patch repair is reduced, given the range of chemically
different carcinogens employed. Whether some of the positive
responses observed in the SSB assays were a direct result of cell
necrosis is discussed below.
Assays for SSB and UDS would be expected to be primarily
sensitive to the 4 confirmed genotoxins represented here, HMPA,
o-toluidine, safrole, and acrylonitrile, and this was observed (cf.
Table 3 and the discussion of the yeast and Drosophila gene-
mutation assays in this section). The only confirmed success with
the remaining 4 carcinogens was the ability of all of the SSB
assays to detect DES. Non-reproducible activity was seen in both
classes of assay for benzene (2/6), benzoin (2/5), and DEHP (1/5).
The latter isolated positive response was strongly endorsed by the
investigator (4.4.3) and requires further investigation.
A topic for future research is the extent to which cell
necrosis, with the consequent formation of broken DNA, contributes
to the outcome of SSB assays. For example, the isolated positive
response observed for benzene (assay 4.3.2) was only observed at
higher dose levels than those employed by the other 2 investigators
in this group. This response may, therefore, represent a technical
false-positive response induced solely by DNA strand breakage as
the result of toxicity. Equally, the extent to which
"mitotic" DNA synthesis has been successfully blocked during the
conduct of a scintillation UDS assay relates directly to the
credibility accorded to weak induced effects. The above 2 concerns
may explain some of the divergencies of data evident within these 3
classes of assay for the present chemicals.
Given the simple requirement to provide evidence of the ability
of a chemical to induce primary DNA damage in mammalian cells, the
investigator is faced with the following decisions:
(a) If the agent is already defined as a bacterial mutagen,
any of the 3 classes of assay (SSB or UDS via
autoradiography or scintillation counting) discussed
herein could be used.
(b) If the agent is inactive as a bacterial mutagen, use of
either the SSB or scintillation counted UDS assays could
present advantages, depending on the chemicals involved.
The results obtained for HMPA and DES suggest that the use
of both assays should be considered, if an assay for "DNA
damage" is required as a complement to bacterial
mutagenicity.
It was, therefore, concluded that the simple legislative
requirement for "evidence of DNA damage" was too vague to be
meaningful. If complementary data are being sought, the
autoradiographic end-point should be avoided and data should be
derived from an SSB assay and/or scintillation counted UDS assay.
The failure of the CSSTT to indicate clearly a single class of
assay, within this group, for routine adoption suggests that
attention should be given to developing the sensitivity of DNA
damage assays to the point that only a single test is required.
The relative performance of these 3 classes of DNA repair assays is
shown in Fig. 3.
9.4. Assays for the Induction of Aneuploidy
In the past, assays for the induction of aneuploidy have not
generally been considered as predictive tests for carcinogenicity.
In the first place, few appropriate in vitro assays existed, and
secondly, the induction of aneuploidy was assumed to have greater
relevance for the promotional, as opposed to the initiatory stages
of carcinogenesis. Thus, while the previous data base has included
the evaluation of a few reference genotoxins such as MNU, it has
mainly focused on cancer-promoting agents such as the phorbol
esters and saccharin, etc. Two investigators have provided the
greater part of the present data base for this class of assay by
presenting data derived from yeast assays (2.4.1.3). In addition,
data derived using mammalian cells were presented (4.9.1) together
with a set of data observed in Aspergillus (2.4.4).
The performance of the aneuploidy assays in the CSSTT was
impressive (Table 2 and 3). In the case of the 2 sets of yeast
data provided from one laboratory (2.4.1 and 2.4.3), a 100%
correlation between the reported carcinogenicity of the test agents
and their activity in vitro was observed. In particular, the
unanimous detection of both DEHP and phenobarbital by these
aneuploidy assays is unique in this study.
The primary conclusion regarding this group of assays is,
therefore, that they present high promise for development as a
class of complementary assays. The development phase should be
influenced by the following considerations, some of which are of
sufficient importance to ensure that they are resolved before any
particular aneuploidy assay is recommended for general adoption.
(a) The findings of the CSSTT suggest that the induction of
aneuploidy may be a critical step in the mechanism of
action of complete carcinogens. These observations
require independent confirmation in other laboratories,
for both the yeast and the mammalian cell systems.
(b) In the case of the D61-M yeast data from the 2
laboratories considered, major discrepancies were evident.
These were all of the same nature and represented the
failure of one laboratory to detect anything beyond
benzene and DES. It is possible that this divergence was
due to technical factors affecting the mitotic or
metabolic activity of the yeast. For example, evidence
from the fungal studies supported an earlier observation
that growing cells are more sensitive to genotoxins than
stationary cells. This could be because of the reduced
metabolic activity of stationary cells. Thus, this
divergence might be due to a relatively minor technical
consideration, but it must be defined, understood,
corrected, and defended in the literature before any
general adoption of yeast aneuploidy assays can be
recommended. The question of whether the D6 strain can be
regarded as repetitive of the D61-M strain also requires
resolution; if it is to be retained, its efficacy must be
confirmed in other laboratories.
(c) The isolated set of Aspergillus data suggests that this
assay is relatively insensitive. Clearly, if the
induction of aneuploidy is to be pursued as a useful
complementary end-point, and if Aspergillus nidulans is to
be selected as the marker organism, a separate data base
will have to be generated to justify this choice.
(d) Requirements similar to those in (c) must apply to the
isolated set of aneuploidy data generated in the Chinese
hamster liver fibroblast assay (4.9.1). This assay is
relatively novel; thus, apart from an expansion of the
data base and the commissioning of independent studies,
the following point requires attention. The cells used in
this study were of between the 9 - 15th passage. The
corollary for using such a spread of cell passages is that
sufficient evidence must be presented on the chromosomal
number and distribution in each cell line before test data
can be interpreted with confidence. Also, the multiple
scoring of hyper, hypo, and polyploid cells presents a
complex data base, the statistical interpretation of which
might not yet have been optimized. Furthermore, the
possible karyotypic instability of higher passage cells
suggests that the current practice of scoring only 200
cells each in test and control cultures might not be
adequate.
The aneuploidy findings of the CSSTT, therefore, provide an
exciting area for future research. However, the role of such
assays as routinely employed complementary tests must await the
acquisition of further data and the outcome of developmental
studies. The related polyploidy assays are considered in section
9.7.
9.5. Mammalian Cell Gene-Mutation Assays
Mammalian cell gene-mutation assays, together with chromosomal-
aberration assays, are high on the list of optional secondary
assays referred to by legislative authorities. The performance of
these assays was, thus, of particular interest. The mammalian
gene-mutation assays constituted the largest group of assays in the
project.
Sixteen sets of data were considered, 6 of which were based on
nominally the same assay (L5178Y), employing the same selective
agent (trifluorothymidine).
Qualitative inspection of the total data base for the mammalian
gene-mutation assays (Table 2), and of that produced following the
primary reduction step (Table 4), was not, as pointed out by the
working group, initially encouraging. The overall sensitivity of
the collected gene-mutation and chromosomal-aberration assays were
similar (Table 3; 62 and 67% and 46 and 53% overall sensitivity,
respectively, for the 2 groups of 4 carcinogens). The value of
such numerical indices can be questioned, but they are supported by
the individual comparisons drawn in Table 3 for the activity of
each chemical in these 2 classes of assay. Thus, similar overall
sensitivity was observed in both classes of assay for most of the
chemicals. Fig. 4 and 5 show the comparative sensitivity
histograms with the Salmonella assays.

 The working-group discussions on the gene-mutation assays
identified a number of specific factors that could have led to the
many disagreements in assay responses and which, if followed up,
should lead to the development of a greatly improved assay system.
Each of these points was concerned with protocol details, all of
which were of relevance to all of these assays, irrespective of the
cell line, selective agent, etc., employed. Some of these
recommendations are of immediate relevance for the interpretation
of weak assay responses (an important matter that is dealt with in
extenso in the full account of the project). The common feature of
the present test chemicals is that they tend to yield weak
responses in vitro. Thus, it is impossible to discern at this
stage which of the many weak positive responses recorded in this
assay group were real and which were illusions created by
inadequate experimentation. Equally, some of the negative
responses observed may have been attributable to experimental
deficiencies rather than to an absolute inadequacy of the assay to
detect the carcinogen. Suggested protocol modifications,
especially those associated with the repetition of experiments and
the use of appropriate cell numbers, should reduce the incidence of
"technical" false-positive and false-negative responses. The data
base that would have been produced in this study, had such
technical modifications already been incorporated, would doubtless
have been much more consistent. However, in addition to possible
protocol deficiencies, which it is anticipated can be remedied,
there remain 2 other important questions.
The first is the extent to which this class of assay can
realistically be reduced to a common cell line using one or more
common selective agents. This would obviously represent a major
advance, as it would enable meaningful interlaboratory comparisons
of data to be made and engender a feeling of confidence that once a
test chemical had been evaluated for mammalian cell gene
mutagenicity, and a conclusion drawn, this would not be reversed on
retesting in another, but similar assay. The need to try to devise
such a system, suitable for general screening was, in fact, the
final recommendation of the assay working group. Some of the
divergencies in test results, evident in this study, were probably
due to the differential sensitivity and metabolic competence of the
several cell lines employed, and this problem cannot be resolved by
adoption of a single common cell line. However, the range of
genetic loci observed and selective agents used probably led to
many instances of true differential sensitivity. The need to use
at least 2 different genetic markers, when a wide range of
genotoxic agents are to be detected has been discussed elsewhere,
and instances of 2 loci being monitored concomitantly have been
reported (Barrett et al., 1981). A considerable amount of
developmental work will be required to discover the degree to which
this class of assay can be standardized with a retention of overall
sensitivity.
The second concern is related to whether these assays are
suitable, by definition, to act as complements to the Salmonella
assays. The results from the yeast and Drosophila assays were not
encouraging and indicated that these particular assays were not
optimal for this purpose, because of the similar nature of the
genetic event being monitored. Similar concerns apply to mammalian
cell gene-mutation assays. Thus, the concept that some agents may
be specifically clastogenic is an argument against the
incorporation of only gene-mutation assays in a battery. Given
that there is usually a requirement in legislative guidelines to
assess a compound for clastogenic activity, the precise role of
mammalian cell gene-mutation assays needs to be reviewed. These
considerations do not necessarily detract from their use as
confirmatory assays. Thus, if it is considered necessary to
evaluate whether a bacterial gene mutagen is similarly active in
mammalian cells, their use is obvious.
The future use of mammalian gene-mutation assays is, therefore,
likely to be influenced by a variety of considerations. The first
and most fundamental is the need for a precise exposition of their
proposed role in the detection of potential carcinogens in vitro.
If they are to be employed only to confirm activity observed in the
Salmonella assay, then some justification should be provided for
the implied existence of a group of established non-carcinogens
that are mutagenic to Salmonella, yet inactive in a well-conducted
mammalian cell gene-mutation assay. If they are to be used as a
complementary assay to the Salmonella assay, urgent attention
should be given to protocol development along the lines outlined in
the assay working group report. The selection of a single assay,
or the development of a single multi-locus assay, may form part of
this development process. However, the wisdom of employing 2 gene-
mutation assays for screening purposes now needs careful
consideration.
Set against these rather demanding requirements are the
following 2 more positive observations. First, it is significant
that mammalian cell gene-mutation assays formed the largest group
of closely-related assays considered in this study. This is
indicative of the current widespread use of these tests and implies
that, in many laboratories, they are performing to an acceptable
standard for reference mutagenic carcinogens such as 2AAF, MNU,
etc. The large number of laboratories carrying out these tests is,
of course, mainly the result of legislative stimulus, but had the
tests proved generally insensitive or unreliable, it would not have
been possible to collect so many together for this study. This is
in contrast to most of the other assays in the study and provides a
strong stimulus for further work to refine them to the point that
they can be used with confidence for the detection of weak
responses. Second, although this study was concerned primarily
with the detection in vitro of potential mammalian carcinogens, the
possible mutagenic properties of chemicals for in vivo somatic and
germ cells are becoming of increasing interest. If the specific
requirement to assay for possible mammalian in vivo mutagens is a
consideration, the role of in vitro gene-mutation assays assumes a
different complexion.
9.6. Chromosomal-Aberration Assays
As indicated above, the performance of this class of assay was
similar to, but slightly better than, that of the mammalian cell
gene-mutation assays. As with the gene-mutation assays,
chromosomal-aberration tests are generally recommended in almost
all legislative guidelines as suitable for use in conjunction with
a bacterial-mutation test, when screening chemicals for new
potential carcinogens. However, while it is currently normal
practice to select the L5178Y cell line when conducting a gene-
mutation experiment, a range of possible cell lines could be chosen
for cytogenetic studies. Thus, in the CSSTT, 4 distinct cell lines
(CHO cells, human lymphocytes, Chinese hamster liver fibroblasts
(CHI-L), and Chinese hamster lung cells (CHL)) were selected, each
of which had been used previously for such studies, and each of
which could be considered suitable for use, when submitting
genotoxicity test data on a new chemical.
A distinct attribute of the cytogenetics assay data was that,
although a range of cytogenetic disturbances can be classified as
chemically-induced damage, the data base was more ordered than
might have been anticipated, because of the common reporting format
employed by most investigators. This was achieved, in advance, by
the assay group Chairman and was based substantially on the
guidelines for assay data reporting suggested in the UKEMS criteria
document (UKEMS, 1983). Further sources of protocol variation can,
therefore, be associated directly with technical decisions made by
individual investigators, including such issues as doselevels,
sampling times, methods of statistical analysis, cell line adopted,
etc.
As would be expected from the similar overall detection rates
of the gene-mutation and chromosomal-aberration assays (Table 3,
and Fig. 4 and 5), simple inspection of the chromosomal-aberration
data base (Table 2 and 4) indicates a disappointing scatter of
positive and negative observations. Certain consistencies were,
however, evident. For example, general agreement was observed in
the 3 cell lines studied for the clastogenicity of both
acrylonitrile (4/4+) and phenobarbital (3+ and 1? of 4). With the
exception of a questionable response in one of the 3 CHO assays
(4.5.3), DEHP gave negative results. Benzoin gave 3 negative
responses and a single, but confirmed, positive response in the CHL
assay. A partially consistent data set was observed for HMPA,
which gave a negative response in each of the 2 CHO assays but was
active in the 3 other cytogenetic assays. This may reflect the
general technical inadequacy of these assay protocols or represent
a general failure of this cell line to metabolize HMPA, a genotoxic
species. One of the 2 CHO SCE assays gave a positive response for
this agent, and this weakens, but does not destroy, the metabolic
deficiency rationale.
Marked divergencies in assay responses were evident for the
remaining 5 chemicals ( o-toluidine, safrole, benzene, caprolactam,
and DES). An approximately equal incidence of positive and
negative responses was observed for these agents and each was
regarded as clastogenic in the assay working-group "consensus"
results ( o-toluidine 3/5 + safrole 2/4 +, benzene 3/6 + caprolactam
2/6 +, and DES 3/4 +). Important protocol deficiencies were
identified in the assay working-group discussions. These
deficiencies were generally associated with the selection of
inappropriate sampling times or dose levels, and if further work
could prove them to be the critical contributors to false-negative
assay responses, then the performance of these assays would be
dramatically improved.
For the purpose of defining a complementary assay within the
present context, the 4 classes of chromosomal-aberration assays
cannot be considered as equivalent. For example, the CHL assay has
been extensively studied and reported on in the literature, but its
use is centred mainly in a single laboratory. Assays involving
human lymphocytes, especially if a rat liver S9 mix is incorporated
into the protocol, are few (Buckton & Evans, 1982). The study data
using Chinese hamster liver fibroblasts formed the greater part of
the accepted data base for this assay. In contrast, the CHO assay
probably represented the one most widely used in industrial and
academic laboratories. This assay can, therefore, be regarded as
particularly relevant for the objectives of the project, as
probably more data are generated using it than any other, when
compounds are assessed prior to legal registration. Given that
some of the negative responses observed in CHO cells might be
corrected by the suggested modifications to the assay protocol,
there is still the problem that, at present, different responses
may be observed in different laboratories using nominally the same
assay. This is similar to the situation currently evident with the
L5178Y gene-mutation assays and, as in that case, urgent attention
to further protocol design and data interpretation is indicated.
The CHL assay (4.5.6), or perhaps the protocol by which it was
conducted, was more sensitive than the others in this group. By
the same criteria, the partial data base evident for the human
lymphocyte assay was also encouraging and worthy of elaboration.
As in other assay systems, it was noted how "assays" were judged,
or misjudged, by the test protocols adopted. This important aspect
of test systems must be resolved, because of the real possibility
that, if more detailed or refined protocols had been used, many
assays would have appeared in a much more favourable light.
An advantage of the chromosomal assays is the extent of the
existing data base and the technical skill that is available for
the conduct of these assays. Thus, it can be expected that
acceptable protocol modifications could be rapidly developed and
disseminated for general adoption. A second advantage is that a
range of supplementary and probably "complementary" end-points can
be easily assessed, once the decision has been taken to conduct
cytogenetic studies within a laboratory. These include the
recording of polyploid cells, an assessment of aneuploidy, and the
studying of sister chromatid exchanges (SCE). Aneuploidy induction
in mammalian cells is considered as a separate assay, and the
induction of SCEs is discussed below. However, the induction of
polyploidy is pertinent to the present discussion as it can be
assessed either concomitantly with, or in parallel to, chromosomal-
aberration assays, depending on the sampling time of the assay.
9.7. Assays for Polyploidy Induction
The chemical induction of polyploidy was discounted as a
separate assay by the working group, but the data generated were
considered because of the interest of the results that were
available. Three complete sets of polyploidy data were presented
(Table 2; assays 4.8.1 (CHL), 4.8.2 (CH liver), and 4.8.3 (RL4, rat
liver)), and each detected DES as positive. This activity is
probably associated with the ability of DES to induce aneuploidy,
spindle effects, and chromosomal aberrations. In the last 2 assays
(4.8.2 and 4.8.3), the remaining 9 compounds were found to be
negative while, in the CHL assay (4.8.1), o-toluidine and
caprolactam were detected as active. These responses indicate that
this genetic end-point is unlikely to produce false-positive
responses; in fact, activity seems to be rare.
The mode of action of DES, benzene, and DEHP as carcinogens is
less clearly defined than that of most other agents. Not only are
they inactive in the Salmonella assay, but they were difficult to
detect as positive in many of the eukaryotic assays used in the
CSSTT. Furthermore, on the basis of their chemical structure and
known or anticipated metabolism, these agents do not appear to be
likely to interact covalently with DNA. It was, therefore,
perceived by many investigators to be of particular interest that
each of these agents was capable of inducing both aneuploidy and
cell transformation in vitro. Furthermore, if "associated"
chromosomal-assay data are considered in conjunction with the
aneuploidy findings, a potentially useful pattern emerges, as shown
in Table 5.
Table 5. Comparative performance of DES, benzene, and DEHP in the classes of
assay showna
----------------------------------------------------------------------------------
Cell Aneuploidy Clastogenicity Polyploidy Spindle studies
transformation (assay 4.9.2)
----------------------------------------------------------------------------------
DES + + + + spindle damage
benzene + + + - modified spindle
function
DEHP + + - - modified spindle
function
----------------------------------------------------------------------------------
a Results are based on the consensus conclusions given by the working groups.
From Table 5, it can be seen that DES was clearly active in
each of the 3 categories of assay, and that its mode of action in
each might be closely related to its ability to damage the
microtubules of the metaphase spindle. In contrast, DEHP was
inactive in the 3 polyploidy assays, as well as in the chromosomal
assays. The question therefore arises of whether the cell-
transforming and aneuploidy-inducing properties of DEHP are
mechanistically unrelated to those of DES. If totally different
mechanisms of action are involved, then the relationship of these
findings to the carcinogenicity of these agents may equally be
different. In this connection, the difference between modification
of spindle integrity and spindle function may be important. Thus,
it is concluded that if an assay for chromosomal aberrations is to
be used as a complementary test, the determination of changes in
ploidy may be a useful additional parameter to measure. In some
assays, it may not be possible to score both chromosomal
aberrations and ploidy, at the same time, mainly because of the
need to progress to the second mitosis after treatment, for
polyploidy to become evident. However, the maintenance of a
parallel culture after treatment for separate assessment of ploidy
should present few difficulties.
9.8. Sister Chromatid Exchange (SCE) Assays
The relationship between the chemical induction of SCEs and the
production of chromosomal aberrations is not clear, and the
mechanism of this phenomenon is obscure. The role of SCE assays in
the detection of carcinogens has yet to be defined. For these
reasons, and because their respective test protocols are quite
distinct, SCE assays have been considered separately from the
aberration assays in the present discussion.
One set of data was omitted from Table 4 owing to the recording
of 6 out of 6 negative responses (4.6.4) and, in the remaining 5
SCE assays, only 12 positive responses were observed out of 41
determinations. These positive responses were not randomly
distributed among the compounds as can be seen in Table 3. Two-
thirds of the SCE assays made on HMPA, o-toluidine, safrole, and
acrylonitrile gave positive responses, while only a single positive
response was observed among the 14 determinations made on benzene,
DEHP, DES, or phenobarbital. This grouping of positive responses
is very similar to that seen with the Drosophila mutation assays
and to a lesser extent by the yeast gene-mutation and UDS
(scintillation) assays. This supports earlier suggestions that the
induction of SCEs is more closely related to gene mutagenicity than
to clastogenicity. No activity was observed for the 2 non-
carcinogens.
Given that the greatest number of positive responses was
observed in this study for the first 4 "genotoxic" carcinogens
shown in Table 4, the SCE findings are unexceptional. The
inability to detect the remaining 4 carcinogens by SCE assays may
provide clues (along with the Drosophila assays) regarding the
mechanism of carcinogenic action of these agents. An observation
of particular interest is that 3 carcinogens, benzene, DES, and
phenobarbital (but not DEHP) were each active in one or more of the
other chromosome-based assays; their inactivity in the SCE assays,
therefore, becomes pointed.
The SCE assays, therefore, appear to be of limited advantage as
complementary assays. Together with several other assays, they
successfully detected the "agreed" genotoxins in the CSSTT and, as
such, the assay seems to be more closely related to the Drosophila
and yeast gene-mutation assays. The low detection rates evident
for the 4 remaining carcinogens suggest that SCE assays are less
generally useful as a complementary test than the chromosomal-
aberration and mammalian cell gene-mutation assays.
9.9. Transformation Assays
The working group, for reasons briefly referred to earlier, did
not consider this class of assay to be suitable for a generally-
applicable complementary assay. However, the good overall
performance of the assay in the CSSTT (Table 3) warrants further
consideration. The chemical induction, in vitro, of the
transformed cell phenotype is appealing, because the derived cell
morphology and behaviour in vivo and in vitro bear a striking
resemblance to the malignant phenotype. The many different
definitions of the transformed cell phenotype (Weinstein et al.,
1976), together with the technical difficulties that accompany the
conduct of such assays have proved to be major factors contributing
to the slow progress that has been made in the general use of these
tests for the detection of possible new carcinogens. The initial
promise shown by the BHK-21 cell-transformation assay was due to
the compatibility of these cells with S9 mix (Styles, 1977), but
progress was impeded by the karyotypic instability of this cell
line and inter-laboratory reproducibility problems encountered at
an early stage (de Serres & Ashby, 1981). The present study
represents the first occasion on which the mouse C3H 10T 1/2 assay
and the Syrian hamster embryo (SHE) assay have been directly
compared in a general study, yet each has an impressive history in
a few laboratories. Although both of these classes of assay
performed well in the CSSTT, there was a marked reluctance among
the assay investigators when it came to suggesting any of these
tests as suitable for routine adoption as a complementary assay.
The insensitivity to the present carcinogens of the Balb C assay
(4.2.1) and the CHO assay (4.2.8) confirmed previous observations
that established cell lines tend to become metabolically
incompetent, yet the metabolically competent primary cell systems
suffer from other problems associated with the criteria used to
select clones of cells for study, and the selection of appropriate
media.
Recent studies associated with the probable role of activation
of the oncogene in the etiology of cancer have invoked the
transformed cell phenotype as a critical marker; thus, the future
of this class of assay in cancer research seems assured.
Nonetheless, the practical problems that would probably be
encountered by a new laboratory endeavouring to establish such an
assay are probably still sufficient to warn against its use as a
routinely-employed complementary test.
10. SELECTION OF A PREFERRED COMPLEMENTARY ASSAY
No single assay was defined by the CSSTT as giving high
carcinogen sensitivity and specificity together with an established
ability to perform optimally in several laboratories. It was
concluded, however, that there was compelling evidence both from
within and beyond the collaborative study, for the adoption of a
chromosomal-aberration assay as the reference complementary test.
This conclusion was drawn on the basis of the following data and
reasoning:
(a) Eight classes of tests, as listed in Table 3, were assessed
for performance as complementary assays, in the foregoing
discussion. Of these, the gene-mutation assays in yeast
and Drosophila were not favoured because of their
repetition of genetic end-point and their poor performance
with the last 4 carcinogens (benzene, DES, phenobarbital,
and DEHP). Assays for aneuploidy were not recommended
because of their novelty, which necessarily prevented
independent studies from being commissioned. The DNA-
repair assays showed poor sensitivity for the last 4
carcinogens, and, occasionally, dramatic differences in
sensitivity between the single-strand breakage and
scintillation UDS tests were evident (Fig. 3). The data
base for the SCE assays suggested marked insensitivity to
the last 4 carcinogens (Fig. 6). Several of the above
assays were, therefore, defined as capable of acting as a
partial complement to the Salmonella assay, but none could
be recommended for routine use as a generally sensitive
complementary test.
The working-group discussions on the gene-mutation assays
identified a number of specific factors that could have led to the
many disagreements in assay responses and which, if followed up,
should lead to the development of a greatly improved assay system.
Each of these points was concerned with protocol details, all of
which were of relevance to all of these assays, irrespective of the
cell line, selective agent, etc., employed. Some of these
recommendations are of immediate relevance for the interpretation
of weak assay responses (an important matter that is dealt with in
extenso in the full account of the project). The common feature of
the present test chemicals is that they tend to yield weak
responses in vitro. Thus, it is impossible to discern at this
stage which of the many weak positive responses recorded in this
assay group were real and which were illusions created by
inadequate experimentation. Equally, some of the negative
responses observed may have been attributable to experimental
deficiencies rather than to an absolute inadequacy of the assay to
detect the carcinogen. Suggested protocol modifications,
especially those associated with the repetition of experiments and
the use of appropriate cell numbers, should reduce the incidence of
"technical" false-positive and false-negative responses. The data
base that would have been produced in this study, had such
technical modifications already been incorporated, would doubtless
have been much more consistent. However, in addition to possible
protocol deficiencies, which it is anticipated can be remedied,
there remain 2 other important questions.
The first is the extent to which this class of assay can
realistically be reduced to a common cell line using one or more
common selective agents. This would obviously represent a major
advance, as it would enable meaningful interlaboratory comparisons
of data to be made and engender a feeling of confidence that once a
test chemical had been evaluated for mammalian cell gene
mutagenicity, and a conclusion drawn, this would not be reversed on
retesting in another, but similar assay. The need to try to devise
such a system, suitable for general screening was, in fact, the
final recommendation of the assay working group. Some of the
divergencies in test results, evident in this study, were probably
due to the differential sensitivity and metabolic competence of the
several cell lines employed, and this problem cannot be resolved by
adoption of a single common cell line. However, the range of
genetic loci observed and selective agents used probably led to
many instances of true differential sensitivity. The need to use
at least 2 different genetic markers, when a wide range of
genotoxic agents are to be detected has been discussed elsewhere,
and instances of 2 loci being monitored concomitantly have been
reported (Barrett et al., 1981). A considerable amount of
developmental work will be required to discover the degree to which
this class of assay can be standardized with a retention of overall
sensitivity.
The second concern is related to whether these assays are
suitable, by definition, to act as complements to the Salmonella
assays. The results from the yeast and Drosophila assays were not
encouraging and indicated that these particular assays were not
optimal for this purpose, because of the similar nature of the
genetic event being monitored. Similar concerns apply to mammalian
cell gene-mutation assays. Thus, the concept that some agents may
be specifically clastogenic is an argument against the
incorporation of only gene-mutation assays in a battery. Given
that there is usually a requirement in legislative guidelines to
assess a compound for clastogenic activity, the precise role of
mammalian cell gene-mutation assays needs to be reviewed. These
considerations do not necessarily detract from their use as
confirmatory assays. Thus, if it is considered necessary to
evaluate whether a bacterial gene mutagen is similarly active in
mammalian cells, their use is obvious.
The future use of mammalian gene-mutation assays is, therefore,
likely to be influenced by a variety of considerations. The first
and most fundamental is the need for a precise exposition of their
proposed role in the detection of potential carcinogens in vitro.
If they are to be employed only to confirm activity observed in the
Salmonella assay, then some justification should be provided for
the implied existence of a group of established non-carcinogens
that are mutagenic to Salmonella, yet inactive in a well-conducted
mammalian cell gene-mutation assay. If they are to be used as a
complementary assay to the Salmonella assay, urgent attention
should be given to protocol development along the lines outlined in
the assay working group report. The selection of a single assay,
or the development of a single multi-locus assay, may form part of
this development process. However, the wisdom of employing 2 gene-
mutation assays for screening purposes now needs careful
consideration.
Set against these rather demanding requirements are the
following 2 more positive observations. First, it is significant
that mammalian cell gene-mutation assays formed the largest group
of closely-related assays considered in this study. This is
indicative of the current widespread use of these tests and implies
that, in many laboratories, they are performing to an acceptable
standard for reference mutagenic carcinogens such as 2AAF, MNU,
etc. The large number of laboratories carrying out these tests is,
of course, mainly the result of legislative stimulus, but had the
tests proved generally insensitive or unreliable, it would not have
been possible to collect so many together for this study. This is
in contrast to most of the other assays in the study and provides a
strong stimulus for further work to refine them to the point that
they can be used with confidence for the detection of weak
responses. Second, although this study was concerned primarily
with the detection in vitro of potential mammalian carcinogens, the
possible mutagenic properties of chemicals for in vivo somatic and
germ cells are becoming of increasing interest. If the specific
requirement to assay for possible mammalian in vivo mutagens is a
consideration, the role of in vitro gene-mutation assays assumes a
different complexion.
9.6. Chromosomal-Aberration Assays
As indicated above, the performance of this class of assay was
similar to, but slightly better than, that of the mammalian cell
gene-mutation assays. As with the gene-mutation assays,
chromosomal-aberration tests are generally recommended in almost
all legislative guidelines as suitable for use in conjunction with
a bacterial-mutation test, when screening chemicals for new
potential carcinogens. However, while it is currently normal
practice to select the L5178Y cell line when conducting a gene-
mutation experiment, a range of possible cell lines could be chosen
for cytogenetic studies. Thus, in the CSSTT, 4 distinct cell lines
(CHO cells, human lymphocytes, Chinese hamster liver fibroblasts
(CHI-L), and Chinese hamster lung cells (CHL)) were selected, each
of which had been used previously for such studies, and each of
which could be considered suitable for use, when submitting
genotoxicity test data on a new chemical.
A distinct attribute of the cytogenetics assay data was that,
although a range of cytogenetic disturbances can be classified as
chemically-induced damage, the data base was more ordered than
might have been anticipated, because of the common reporting format
employed by most investigators. This was achieved, in advance, by
the assay group Chairman and was based substantially on the
guidelines for assay data reporting suggested in the UKEMS criteria
document (UKEMS, 1983). Further sources of protocol variation can,
therefore, be associated directly with technical decisions made by
individual investigators, including such issues as doselevels,
sampling times, methods of statistical analysis, cell line adopted,
etc.
As would be expected from the similar overall detection rates
of the gene-mutation and chromosomal-aberration assays (Table 3,
and Fig. 4 and 5), simple inspection of the chromosomal-aberration
data base (Table 2 and 4) indicates a disappointing scatter of
positive and negative observations. Certain consistencies were,
however, evident. For example, general agreement was observed in
the 3 cell lines studied for the clastogenicity of both
acrylonitrile (4/4+) and phenobarbital (3+ and 1? of 4). With the
exception of a questionable response in one of the 3 CHO assays
(4.5.3), DEHP gave negative results. Benzoin gave 3 negative
responses and a single, but confirmed, positive response in the CHL
assay. A partially consistent data set was observed for HMPA,
which gave a negative response in each of the 2 CHO assays but was
active in the 3 other cytogenetic assays. This may reflect the
general technical inadequacy of these assay protocols or represent
a general failure of this cell line to metabolize HMPA, a genotoxic
species. One of the 2 CHO SCE assays gave a positive response for
this agent, and this weakens, but does not destroy, the metabolic
deficiency rationale.
Marked divergencies in assay responses were evident for the
remaining 5 chemicals ( o-toluidine, safrole, benzene, caprolactam,
and DES). An approximately equal incidence of positive and
negative responses was observed for these agents and each was
regarded as clastogenic in the assay working-group "consensus"
results ( o-toluidine 3/5 + safrole 2/4 +, benzene 3/6 + caprolactam
2/6 +, and DES 3/4 +). Important protocol deficiencies were
identified in the assay working-group discussions. These
deficiencies were generally associated with the selection of
inappropriate sampling times or dose levels, and if further work
could prove them to be the critical contributors to false-negative
assay responses, then the performance of these assays would be
dramatically improved.
For the purpose of defining a complementary assay within the
present context, the 4 classes of chromosomal-aberration assays
cannot be considered as equivalent. For example, the CHL assay has
been extensively studied and reported on in the literature, but its
use is centred mainly in a single laboratory. Assays involving
human lymphocytes, especially if a rat liver S9 mix is incorporated
into the protocol, are few (Buckton & Evans, 1982). The study data
using Chinese hamster liver fibroblasts formed the greater part of
the accepted data base for this assay. In contrast, the CHO assay
probably represented the one most widely used in industrial and
academic laboratories. This assay can, therefore, be regarded as
particularly relevant for the objectives of the project, as
probably more data are generated using it than any other, when
compounds are assessed prior to legal registration. Given that
some of the negative responses observed in CHO cells might be
corrected by the suggested modifications to the assay protocol,
there is still the problem that, at present, different responses
may be observed in different laboratories using nominally the same
assay. This is similar to the situation currently evident with the
L5178Y gene-mutation assays and, as in that case, urgent attention
to further protocol design and data interpretation is indicated.
The CHL assay (4.5.6), or perhaps the protocol by which it was
conducted, was more sensitive than the others in this group. By
the same criteria, the partial data base evident for the human
lymphocyte assay was also encouraging and worthy of elaboration.
As in other assay systems, it was noted how "assays" were judged,
or misjudged, by the test protocols adopted. This important aspect
of test systems must be resolved, because of the real possibility
that, if more detailed or refined protocols had been used, many
assays would have appeared in a much more favourable light.
An advantage of the chromosomal assays is the extent of the
existing data base and the technical skill that is available for
the conduct of these assays. Thus, it can be expected that
acceptable protocol modifications could be rapidly developed and
disseminated for general adoption. A second advantage is that a
range of supplementary and probably "complementary" end-points can
be easily assessed, once the decision has been taken to conduct
cytogenetic studies within a laboratory. These include the
recording of polyploid cells, an assessment of aneuploidy, and the
studying of sister chromatid exchanges (SCE). Aneuploidy induction
in mammalian cells is considered as a separate assay, and the
induction of SCEs is discussed below. However, the induction of
polyploidy is pertinent to the present discussion as it can be
assessed either concomitantly with, or in parallel to, chromosomal-
aberration assays, depending on the sampling time of the assay.
9.7. Assays for Polyploidy Induction
The chemical induction of polyploidy was discounted as a
separate assay by the working group, but the data generated were
considered because of the interest of the results that were
available. Three complete sets of polyploidy data were presented
(Table 2; assays 4.8.1 (CHL), 4.8.2 (CH liver), and 4.8.3 (RL4, rat
liver)), and each detected DES as positive. This activity is
probably associated with the ability of DES to induce aneuploidy,
spindle effects, and chromosomal aberrations. In the last 2 assays
(4.8.2 and 4.8.3), the remaining 9 compounds were found to be
negative while, in the CHL assay (4.8.1), o-toluidine and
caprolactam were detected as active. These responses indicate that
this genetic end-point is unlikely to produce false-positive
responses; in fact, activity seems to be rare.
The mode of action of DES, benzene, and DEHP as carcinogens is
less clearly defined than that of most other agents. Not only are
they inactive in the Salmonella assay, but they were difficult to
detect as positive in many of the eukaryotic assays used in the
CSSTT. Furthermore, on the basis of their chemical structure and
known or anticipated metabolism, these agents do not appear to be
likely to interact covalently with DNA. It was, therefore,
perceived by many investigators to be of particular interest that
each of these agents was capable of inducing both aneuploidy and
cell transformation in vitro. Furthermore, if "associated"
chromosomal-assay data are considered in conjunction with the
aneuploidy findings, a potentially useful pattern emerges, as shown
in Table 5.
Table 5. Comparative performance of DES, benzene, and DEHP in the classes of
assay showna
----------------------------------------------------------------------------------
Cell Aneuploidy Clastogenicity Polyploidy Spindle studies
transformation (assay 4.9.2)
----------------------------------------------------------------------------------
DES + + + + spindle damage
benzene + + + - modified spindle
function
DEHP + + - - modified spindle
function
----------------------------------------------------------------------------------
a Results are based on the consensus conclusions given by the working groups.
From Table 5, it can be seen that DES was clearly active in
each of the 3 categories of assay, and that its mode of action in
each might be closely related to its ability to damage the
microtubules of the metaphase spindle. In contrast, DEHP was
inactive in the 3 polyploidy assays, as well as in the chromosomal
assays. The question therefore arises of whether the cell-
transforming and aneuploidy-inducing properties of DEHP are
mechanistically unrelated to those of DES. If totally different
mechanisms of action are involved, then the relationship of these
findings to the carcinogenicity of these agents may equally be
different. In this connection, the difference between modification
of spindle integrity and spindle function may be important. Thus,
it is concluded that if an assay for chromosomal aberrations is to
be used as a complementary test, the determination of changes in
ploidy may be a useful additional parameter to measure. In some
assays, it may not be possible to score both chromosomal
aberrations and ploidy, at the same time, mainly because of the
need to progress to the second mitosis after treatment, for
polyploidy to become evident. However, the maintenance of a
parallel culture after treatment for separate assessment of ploidy
should present few difficulties.
9.8. Sister Chromatid Exchange (SCE) Assays
The relationship between the chemical induction of SCEs and the
production of chromosomal aberrations is not clear, and the
mechanism of this phenomenon is obscure. The role of SCE assays in
the detection of carcinogens has yet to be defined. For these
reasons, and because their respective test protocols are quite
distinct, SCE assays have been considered separately from the
aberration assays in the present discussion.
One set of data was omitted from Table 4 owing to the recording
of 6 out of 6 negative responses (4.6.4) and, in the remaining 5
SCE assays, only 12 positive responses were observed out of 41
determinations. These positive responses were not randomly
distributed among the compounds as can be seen in Table 3. Two-
thirds of the SCE assays made on HMPA, o-toluidine, safrole, and
acrylonitrile gave positive responses, while only a single positive
response was observed among the 14 determinations made on benzene,
DEHP, DES, or phenobarbital. This grouping of positive responses
is very similar to that seen with the Drosophila mutation assays
and to a lesser extent by the yeast gene-mutation and UDS
(scintillation) assays. This supports earlier suggestions that the
induction of SCEs is more closely related to gene mutagenicity than
to clastogenicity. No activity was observed for the 2 non-
carcinogens.
Given that the greatest number of positive responses was
observed in this study for the first 4 "genotoxic" carcinogens
shown in Table 4, the SCE findings are unexceptional. The
inability to detect the remaining 4 carcinogens by SCE assays may
provide clues (along with the Drosophila assays) regarding the
mechanism of carcinogenic action of these agents. An observation
of particular interest is that 3 carcinogens, benzene, DES, and
phenobarbital (but not DEHP) were each active in one or more of the
other chromosome-based assays; their inactivity in the SCE assays,
therefore, becomes pointed.
The SCE assays, therefore, appear to be of limited advantage as
complementary assays. Together with several other assays, they
successfully detected the "agreed" genotoxins in the CSSTT and, as
such, the assay seems to be more closely related to the Drosophila
and yeast gene-mutation assays. The low detection rates evident
for the 4 remaining carcinogens suggest that SCE assays are less
generally useful as a complementary test than the chromosomal-
aberration and mammalian cell gene-mutation assays.
9.9. Transformation Assays
The working group, for reasons briefly referred to earlier, did
not consider this class of assay to be suitable for a generally-
applicable complementary assay. However, the good overall
performance of the assay in the CSSTT (Table 3) warrants further
consideration. The chemical induction, in vitro, of the
transformed cell phenotype is appealing, because the derived cell
morphology and behaviour in vivo and in vitro bear a striking
resemblance to the malignant phenotype. The many different
definitions of the transformed cell phenotype (Weinstein et al.,
1976), together with the technical difficulties that accompany the
conduct of such assays have proved to be major factors contributing
to the slow progress that has been made in the general use of these
tests for the detection of possible new carcinogens. The initial
promise shown by the BHK-21 cell-transformation assay was due to
the compatibility of these cells with S9 mix (Styles, 1977), but
progress was impeded by the karyotypic instability of this cell
line and inter-laboratory reproducibility problems encountered at
an early stage (de Serres & Ashby, 1981). The present study
represents the first occasion on which the mouse C3H 10T 1/2 assay
and the Syrian hamster embryo (SHE) assay have been directly
compared in a general study, yet each has an impressive history in
a few laboratories. Although both of these classes of assay
performed well in the CSSTT, there was a marked reluctance among
the assay investigators when it came to suggesting any of these
tests as suitable for routine adoption as a complementary assay.
The insensitivity to the present carcinogens of the Balb C assay
(4.2.1) and the CHO assay (4.2.8) confirmed previous observations
that established cell lines tend to become metabolically
incompetent, yet the metabolically competent primary cell systems
suffer from other problems associated with the criteria used to
select clones of cells for study, and the selection of appropriate
media.
Recent studies associated with the probable role of activation
of the oncogene in the etiology of cancer have invoked the
transformed cell phenotype as a critical marker; thus, the future
of this class of assay in cancer research seems assured.
Nonetheless, the practical problems that would probably be
encountered by a new laboratory endeavouring to establish such an
assay are probably still sufficient to warn against its use as a
routinely-employed complementary test.
10. SELECTION OF A PREFERRED COMPLEMENTARY ASSAY
No single assay was defined by the CSSTT as giving high
carcinogen sensitivity and specificity together with an established
ability to perform optimally in several laboratories. It was
concluded, however, that there was compelling evidence both from
within and beyond the collaborative study, for the adoption of a
chromosomal-aberration assay as the reference complementary test.
This conclusion was drawn on the basis of the following data and
reasoning:
(a) Eight classes of tests, as listed in Table 3, were assessed
for performance as complementary assays, in the foregoing
discussion. Of these, the gene-mutation assays in yeast
and Drosophila were not favoured because of their
repetition of genetic end-point and their poor performance
with the last 4 carcinogens (benzene, DES, phenobarbital,
and DEHP). Assays for aneuploidy were not recommended
because of their novelty, which necessarily prevented
independent studies from being commissioned. The DNA-
repair assays showed poor sensitivity for the last 4
carcinogens, and, occasionally, dramatic differences in
sensitivity between the single-strand breakage and
scintillation UDS tests were evident (Fig. 3). The data
base for the SCE assays suggested marked insensitivity to
the last 4 carcinogens (Fig. 6). Several of the above
assays were, therefore, defined as capable of acting as a
partial complement to the Salmonella assay, but none could
be recommended for routine use as a generally sensitive
complementary test.
 (b) The transformation working group recommended that assays
for cell transformation were not suitable for routine
adoption because of the technical difficulties of carrying
them out and the remaining uncertainty regarding the
definition of the transformed phenotype. Nonetheless, 6 of
these assays showed high sensitivity to the present 8
carcinogens; in fact, as a class, they performed better
than any other category of test (Table 3). It is clear
that both the C3H 10T 1/2 and the SHE assay are sensitive
assays, and each performed similarly, in the CSSTT, in 2
different laboratories. Nonetheless, the advice of the
assay group is endorsed here; namely, that transformation
assays are not to be recommended for routine adoption, at
present, especially by laboratories new to such techniques.
(c) Two classes of assay remain for consideration, each of
which was well represented in the CSSTT, i.e., mammalian
cell gene-mutation and chromosomal-aberration assays. Both
of these classes of assay showed a similar level of
carcinogen sensitivity, when considering the reduced data
base (Table 3 and 4).
The apparently similar performance of these 2 classes of
assay as complementary tests is shown in Fig. 4 and 5. The
performance of the Drosophila and SCE assays is shown in
Fig. 6 for purposes of comparison and to illustrate that
some assays only partially complement the Salmonella assay.
When considering the total data base (Table 2) for the
gene-mutation and chromosomal-aberration assays, the latter
appear to be more sensitive, yielding 22 positive responses
out of 43 determinations made on the 8 carcinogens (51%),
as opposed to only 40 out of 103 for the mutation assays
(39%). The low sensitivity of some of the mutation assays
led to 6 of the 16 being eliminated in Table 3 as opposed
to only 1 of the 7 cytogenetic assays. The normalized (%)
performances of these 2 classes of assays as complementary
tests are compared in Fig. 7. Similar efficiencies are
evident with the chromosomal assays proving marginally more
sensitive to the carcinogens.
Figs. 4, 5, and 7 enable the problem of reproducibility of
response to be visualized and it is clear that harmonization and/or
extension of assay protocols is indicated for both classes of test.
The effort required to produce a common and sensitive assay
protocol, combined with a realistic assessment of whether such
efforts are likely to be rewarded, constitutes the final factor in
the recommendation of the chromosomal-aberration assays. Four
points appear to be particularly relevant:
(a) chromosomal-aberration assays offer an independent genetic
end-point from the Salmonella assay;
(b) chromosomal-aberration assays have a more extensive
history than the mammalian cell gene-mutation assays; this
means that more laboratories have used them for a longer
period; thus, any protocol corrections required should be
more clearly perceived and easier to institute;
(c) the recommendations for assay protocol modification made
by the chromosomal-aberration working group were more
clearly defined, less fundamental, and seemingly easier to
institute than those made by the gene-mutation working
group; and
(d) once the decision is made to conduct a chromosomal-
aberration assay in a laboratory, the expertise and
facilities required will serve equally for the assessment
of aneuploidy, polyploidy, and SCEs.
(b) The transformation working group recommended that assays
for cell transformation were not suitable for routine
adoption because of the technical difficulties of carrying
them out and the remaining uncertainty regarding the
definition of the transformed phenotype. Nonetheless, 6 of
these assays showed high sensitivity to the present 8
carcinogens; in fact, as a class, they performed better
than any other category of test (Table 3). It is clear
that both the C3H 10T 1/2 and the SHE assay are sensitive
assays, and each performed similarly, in the CSSTT, in 2
different laboratories. Nonetheless, the advice of the
assay group is endorsed here; namely, that transformation
assays are not to be recommended for routine adoption, at
present, especially by laboratories new to such techniques.
(c) Two classes of assay remain for consideration, each of
which was well represented in the CSSTT, i.e., mammalian
cell gene-mutation and chromosomal-aberration assays. Both
of these classes of assay showed a similar level of
carcinogen sensitivity, when considering the reduced data
base (Table 3 and 4).
The apparently similar performance of these 2 classes of
assay as complementary tests is shown in Fig. 4 and 5. The
performance of the Drosophila and SCE assays is shown in
Fig. 6 for purposes of comparison and to illustrate that
some assays only partially complement the Salmonella assay.
When considering the total data base (Table 2) for the
gene-mutation and chromosomal-aberration assays, the latter
appear to be more sensitive, yielding 22 positive responses
out of 43 determinations made on the 8 carcinogens (51%),
as opposed to only 40 out of 103 for the mutation assays
(39%). The low sensitivity of some of the mutation assays
led to 6 of the 16 being eliminated in Table 3 as opposed
to only 1 of the 7 cytogenetic assays. The normalized (%)
performances of these 2 classes of assays as complementary
tests are compared in Fig. 7. Similar efficiencies are
evident with the chromosomal assays proving marginally more
sensitive to the carcinogens.
Figs. 4, 5, and 7 enable the problem of reproducibility of
response to be visualized and it is clear that harmonization and/or
extension of assay protocols is indicated for both classes of test.
The effort required to produce a common and sensitive assay
protocol, combined with a realistic assessment of whether such
efforts are likely to be rewarded, constitutes the final factor in
the recommendation of the chromosomal-aberration assays. Four
points appear to be particularly relevant:
(a) chromosomal-aberration assays offer an independent genetic
end-point from the Salmonella assay;
(b) chromosomal-aberration assays have a more extensive
history than the mammalian cell gene-mutation assays; this
means that more laboratories have used them for a longer
period; thus, any protocol corrections required should be
more clearly perceived and easier to institute;
(c) the recommendations for assay protocol modification made
by the chromosomal-aberration working group were more
clearly defined, less fundamental, and seemingly easier to
institute than those made by the gene-mutation working
group; and
(d) once the decision is made to conduct a chromosomal-
aberration assay in a laboratory, the expertise and
facilities required will serve equally for the assessment
of aneuploidy, polyploidy, and SCEs.
 It is therefore recommended that major efforts be focused on
designing a minimum effective protocol according to which
chromosomal-aberration assays should be conducted, and that this
could constitute a complementary assay for the Salmonella test.
Recourse to other assays may be indicated, in order to assess fully
the in vitro activity of certain test chemicals. Acquisition of
test data from a minimum of these 2 standard assays, if optimally
performed, will enable a greater degree of comparability of data
between both independent laboratories and different countries, and
this can only help the efficient detection of possible new human
carcinogens.
In many testing schemes, resources are spread across a large
number of in vitro assays, and this may lead to assays on a new
chemical being conducted suboptimally. The alternative suggested
here is that with 2 well-conducted and common assays, there could
be enhanced comparability and reliability of data, together with an
increase in carcinogen sensitivity. In addition, the Salmonella
assay should be conducted according to what is now regarded as the
best current practice, i.e., employing strains TA1535, 1537, 1538,
98, and 100 in the presence and absence of an induced rat liver S9
mix (10%) (Ames et al., 1975; UKEMS 1983) and also using a
preincubation protocol with strains TA98 and 100 in the case of
initial negative responses. Use of a different source of S9 mix
(e.g., hamster) and additional levels of rat liver S9 mix (e.g., 4%
and 30%) should also be considered (Venitt et al., 1984). In the
case of volatile chemicals, an enclosed test system should be
employed.
The particular cell line employed for cytogenetic assays may be
critical but should not necessarily be restricted to the widely-
used Chinese hamster ovary (CHO) line. Good results were obtained
with the Chinese hamster lung (CHL) line in the CSSTT, but it is
not commercially available at present, and while human lymphocytes
present a sensitive and readily-available source of primary euploid
cells, further studies will be required to demonstrate the general
usefulness of this system (Buckton & Evans, 1982). The Chinese
hamster liver system protocol was designed to measure both
aberrations and aneuploidy and, likewise, will require further
independent studies to confirm the promise shown in the CSSTT.
11. CONCLUSIONS
The dilemma that preceded the formalization of final
conclusions was difficult to resolve. Several carcinogens of
uncertain genotoxicity were selected for the CSSTT, in order to
ascertain which among the many available eukaryotic in vitro
assays for carcinogens would be most efficient and best suited for
adoption as a complement to the generally available Salmonella
mutation assay, by laboratories engaged in the routine screening of
chemicals for potential carcinogens. The organizing committee was
aware of the impact that any conclusions derived from this study
might have on legislative guidelines, but was primarily interested
in the optimal use of the resources of laboratories that may be
attempting to identify possible new human carcinogens. In order to
provide a comprehensive study, the number of types of assays
included was not limited. This non-restrictive method of
collecting assays presented a unique problem; by implication, some
investigators had been invited to submit their assay for scrutiny
for a purpose other than they had originally intended. For
example, many oncologists conduct cell-transformation, aneuploidy,
UDS, etc., assays for research purposes, in order to study the
relationship between genetic end-points and the subtle nuances of
the cancer process. Such research studies may have supported the
selective use of a particular assay that was not claimed to be of
general usefulness for the present purposes, and which was not
being recommended as such by the investigators. It is, therefore,
important to emphasize that the failure of an assay to qualify as a
complementary assay does not preclude its use for research into the
intricate processes of cancer initiation/promotion. This caution
is most important to remember when whole classes of assays such as
"cell transformation", "metabolic cooperation", etc., have been
dismissed from consideration in this collaborative study. On the
one hand, is the immediate and worldwide need for efficient and
reliable means for screening new chemicals, leading to their
registration as safe new products (Ashby et al., 1983); on the
other, the need to use selected in vitro assays to probe the
etiology of chemically-induced cancer. The present study is aimed
solely at the former endeavour, but its findings might expedite the
achievement of the latter. Ancillary findings of the CSSTT are
first, that certain classes of in vitro assays appear to be
insensitive to the cancer-related activities of certain chemical
carcinogens. Second, it is suggested that a clear distinction
should be made between the use of eukaryotic in vitro assays to
confirm genotoxic activity, seen previously for a chemical in
bacteria, and their use to complement bacterial-mutation assays,
i.e., to detect genotoxins found inactive in bacteria; the choice
of assay and the interpretation of data are influenced by the need
originally perceived.
1. Significant differences exist among individual investigators
conducting nominally identical assays. A direct result of the
CSSTT is that many recommended protocol changes were identified
for all assays and some have already been instituted by a
number of investigators. Inadequate test protocols clearly
contributed to the fact that some assays appeared to be
insensitive. It is likely that the need, currently perceived,
to conduct a range of genotoxicity assays on a new chemical has
been stimulated, in part, by the inadequacy of some test
protocols.
2. Urgent attention should be given to the definition of criteria
for judging positive and negative test responses in all assays.
Methods by which weak responses can be interpreted with
confidence were identified, including the adoption of
appropriately sensitive test protocols, the acquisition of a
sufficiently large control data base, and the use of
appropriate statistical methods for data assessment.
3. Carcinogens that are inactive or difficult to detect in the
Salmonella assay fall into 2 distinct groups in this study.
The first include genotoxins that are probably only non-
mutagenic to Salmonella because of protocol deficiencies
associated with the overall metabolic capacity of the assay
system. These agents, HMPA, o-toluidine, safrole, and
acrylonitrile, were detected in most of the eukaryotic assays
studied. Thus, a range of assays can act as limited
complements to the Salmonella mutation assay. The other group
of carcinogens, benzene, DEHP, DES, and phenobarbital,
possessed a more selective range of genotoxic activities, and
none of the assays was selectively sensitive to them. Of the
assays studied, only the induction of chromosomal aberrations,
transformation, gene mutations in mammalian cells, and
aneuploidy in yeast gave encouraging overall performances for
the 8 carcinogens (Conclusion 9).
4. A surprising finding of the CSSTT was the apparent mutagenicity
observed for the 3 carcinogens, benzene, DES, and phenobarbital
in some of the mammalian cell gene-mutation assays. These
activities were not seen by the majority of investigators in
this assay group and were not generally confirmed in either the
Drosophila or fungal gene-mutation assays. If they prove
repeatable, 3 cell-specific gene mutagens will have been
defined. If they are not repeatable, this will emphasize the
need, already perceived by the assay working group, to improve
the protocol and data assessment processes for this class of
assay. The concomitant monitoring of 2 gene loci in a single-
cell line, and the development of appropriate and common data
assessment procedures represent 2 areas for research with
mammalian cell gene-mutation assays. The precise role of these
assays in routine screening programmes requires clarification;
in particular, whether they are to be used to anticipate
possible mammalian carcinogens or possible mammalian mutagens,
or both. The use of negative results in a mammalian cell gene-
mutation assay to negate positive observations made in the
Salmonella assay may be premature.
5. A range of cell-transformation assays was studied, and the
overall performance of several of them was impressive. The
duplicated studies with the Syrian hamster embryo (SHE) assay
and the mouse C3H10T 1/2 assay gave particularly encouraging
results. Nonetheless, these assays are not recommended for
routine screening purposes, at present, because of the
expertise required to carry them out and because of the
problems that remain with clone selection and judgement of when
chemically-induced transformation has occurred.
6. Both of the non-carcinogens showed low incidences of activity
in vitro, several of which were confirmed and most of which
were weak. The clastogenicity of caprolactam for human
lymphocytes and its somatic cell mutagenicity in Drosophila
were particularly clear-cut. The activities observed for these
agents in certain in vitro assays do not, therefore, correlate
with their rodent carcinogenicity, and thus, they represent
false-positive predictions of carcinogenicity. This underlines
the fact that in vitro assays are of use for predicting, rather
than defining carcinogenic activity.
7. Three of the carcinogens, DES, phenobarbital, and DEHP, were
chosen to evaluate the hypothesis that some chemicals can
induce tumours in rodents without first modifying the integrity
of nuclear DNA. Although these chemicals were generally less
active in the tests than the other carcinogens, they each
displayed a range of genotoxic activities in vitro. DEHP
showed least activity, with levels similar to those observed
for the 2 non-carcinogens. Nevertheless, it induced
significant aneuploidy and cell-transformation effects in an
almost total absence of evidence of a direct interaction with
DNA. The data generated for DES confirmed earlier reports that
it is an aneuploidy-inducing and cell-transforming agent, and
the present study also revealed that it was capable of causing
primary damage to DNA; it cannot, therefore, be regarded as
non-genotoxic. Phenobarbital was perhaps the previously best
documented non-genotoxin of those studied, but it has been
defined herein as a gene mutagen, a clastogen, and an
aneuploidy-inducing agent. It is concluded that the phrase
"non-genotoxic" should only be used when a sufficiently large
genotoxicity data base has been collected.
8. Many of the assays evaluated may have an important research
role. However, if their main function is to act as
confirmatory or complementary assays in conjunction with the
Salmonella test, then substantial redundancy among in vitro
eukaryotic genotoxicity assays is evident.
9. In relation to the stated aim of the CSSTT, it is recommended
that resources be applied to the definition of a generally
acceptable and applicable protocol for the conduct of
chromosomal-aberration assays, preferably in an agreed cell
type. Use of this class of assay in conjunction with an
adequate assessment of the mutagenicity of a chemical for
Salmonella should provide an efficient primary screen for
possible new carcinogens. Certain chemicals that induce
tumours in rodents at elevated dose levels will remain
undetected by these tests, as exemplified by DEHP in the
present study. The "detection", in vitro, of such agents may
be achieved by the incorporation of additional assays, but the
price in terms of a reduction in overall specificity and the
increased burden on technical resources may be high.
Consideration of whether the joint inactivity of a chemical in
an adequate Salmonella assay and an adequate in vitro
cytogenetic assay provides evidence of its inactivity with DNA
and, thus, its probable non-carcinogenicity for mammals, at low
dose levels, is indicated. The adoption of a chromosomal-
aberration assay as a common complementary test would have the
additional advantages of allowing easy comparison of data,
ready access to supplementary cytogenetic assays, and the
provision of test data derived using a genetic end-point,
independent of that of the Salmonella (gene-mutation) assay.
REFERENCES
AMES, B.N., MCCANN, J., & YAMASAKI, E. (1975) Methods for
detecting carcinogens and mutagens with the salmonella/mammalian-
microsome mutagenicity test. Mutat. Res., 31: 347-364.
ASHBY, J. (1983) The distinction between genetic and non-genetic
mechanisms in chemical carcinogenesis. In: Zbinden, G., Cohadon,
F., Detaille, J.Y., & Mazué, G., ed. Current problems in drug
toxicity, London, Paris, John Libby Eurotext.
ASHBY, J. & LEFEVRE, P.A. (1983) Genetic toxicology with
formaldehyde and closely related chemicals. In: Gibson, J.E., ed.
Formaldehyde toxicity, Hemisphere Pub.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr, MATTER,
B.E., & SHELBY, M. (1983) The two IPCS collaborative studies on
short-term tests for genotoxicity and carcinogenicity. Mutat. Res.,
109: 123-126.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr,
MARGOLIN, B.H., MATTER, B.E., & SHELBY, M., ed. (1985) Evaluation
of short-term tests for carcinogens. Report of the International
Programme on Chemical Safety Collaborative Study on in vitro
assays, Amsterdam, Elsevier Science Publishers (Progress in
Mutation Research, Vol. 5).
ATHANASIOU, K. & KRYRTOPOULOS, S.A. (1981) Induction of SCE by
non-mutagenic carcinogens. NATO Adv. Study Inst. Ser A, 40: 557-562
(Chem. abst., 97 18839 (1982).
BARRETT, J.C., WONG, A., & MCLACHLAN, J.A. (1981)
Diethylstilboestrol induced neoplastic transformation without
measurable gene mutation at two loci. Science, 212: 1402-1404.
BUCKTON, K.E. & EVANS, H.J. (1982) Human peripheral blood
lymphocyte cultures. An in vitro assay for the cytogenetic effects
of environmental mutagens. In: Hsu, T.C., ed. Cytogenetic assays of
environmental mutagens, Osman, New Jersey, Allanheld.
CAIRNS, J. (1981) The origin of human cancers. Nature (Lond.),
289: 353-357.
CLEMMENSEN, J. & HJALGRIM-JENSEN, S. (1977) On the absence of
carcinogenicity to man of phenobarbital. In: Statistical studies in
malignant neoplasms, Vol. 5, pp. 38-50.
DEAN, B.J. (1978) Genetic toxicology of benzene, toluene,
xylenes, and phenols. Mutat. Res., 47: 75-97.
DE SERRES, F.J. & ASHBY, J., ed. (1981) Evaluation of short-term
tests for carcinogens: report of the international collaborative
program, North-Holland, New York, Elsevier.
GOLDMACHER, V.S. & THILLY, W.G. (1983) Formaldehyde is mutagenic
for cultured human cells. Mutat. Res., 116: 417-422.
HUFF, J.E. (1982) Caprolactam: condensation of the carcinogenesis
bioassay technical report. Environ. Health Perspect., 45: 199-200.
KLEIN, G. (1981) The role of gene dosage and genetic
transpositions in carcinogenesis. Nature (Lond.), 294: 313-318.
MILVY, P. & WOLFF, M. (1977) Mutagenic studies with
acrylonitrile. Mutat. Res., 48: 217-278.
MOODY, D.E. & REDDY, J.K. (1978) Hepatic peroxisome (microbody)
proliferation in rats fed plasticizers and related compounds.
Toxicol. appl. Pharmacol., 45: 497-504.
PHILLIPS, B.J., JAMES, T.E.B., & GANGOLLI, S.D. (1982)
Genotoxicity studies of di(2-ethylhexyl)phthalate and its
metabolities in CHO cells. Mutat. Res., 102: 297-304.
REDDY, E.P., REYNOLDS, R.K., SANTOS, E., & BARBACID, M. (1982) A
point of mutation is responsible for the acquisition of
transforming properties by the T24 human bladder carcinoma
oncogene. Nature (Lond.), 300: 149-152.
STYLES, J.A (1977) A method for detecting carcinogenic organic
chemicals using mammalian cells in culture. Br. J. Cancer, 36: 558-563.
TABIN, C.J., BRADLEY, S.M., BARGMANN, C.I., WEINBERG, R.A.,
PAPAGEORGE, A.G., SCOLNICK, E.M., DHAR, R., LOWY, D.R., & CHANG,
E.H. (1982) Mechanism of activation of a human oncogene. Nature
(Lond.), 300: 143-149.
TOMITA, I., NAKAMURA, Y., AOKI, N., & INUI, N. (1982)
Mutagenic/carcinogenic potential of DEHP and MEHP. Environ. Health
Perspect., 45: 119-125.
UKEMS (1983) Report of the UKEMS sub-committee on guidelines for
mutagenicity testing, Part I, The United Kingdom Environmental
Mutagen Society.
UNITED STATES PUBLIC LAW (1958) United States Code, title 21,
section 348(c)(3)(A).
US NTP (1980) Benzoin - NTP technical report on the toxicology
and carcinogencity of benzoin, Research Triangle Park, North
Carolina, National Toxicology Program (TR No. 204).
US NTP (1982) Caprolactam - NTP technical report on the
toxicology and carcinogenicity of benzoin, Research Triangle Park,
North Carolina, National Toxicology Program (TR No. 214).
VENITT, S., BUSHELL, C.T., & OSBORNE, M. (1977) Mutagenicity of
acrylonitrile (cyanoethylene) in Escherichia coli. Mutat. Res., 45:
283-288.
VENITT, S., CROFTON-SLEIGH, C., & OSBORNE, M.B. (1984) The hair
dye reagent 2-(2,4-diaminophenoxy) ethanol is mutagenic to
Salmonella typhimurium. Mutat. Res., 135: 31-47.
WEINSTEIN, I.B., WIGLER, M., & STADLER, U. (1976) Analysis of the
mechanism of chemical carcinogenesis of epithelial cell cultures,
Lyons, International Agency for Research on Cancer, (IARC
Scientific Publication No. 12, Screening tests in Chemical
Carcinogenesis).
WEISS, R. (1982) The myconcogene in man and birds. Nature
(Lond.), 299: 9-10.
WILLIAMS, G.M. (1981) Liver carcinogenesis: the role for some
chemicals of an epigenetic mechanism of liver-tumour promotion
involving modification of the cell membrane. Food Cosmet. Toxicol.,
19: 577-583.
It is therefore recommended that major efforts be focused on
designing a minimum effective protocol according to which
chromosomal-aberration assays should be conducted, and that this
could constitute a complementary assay for the Salmonella test.
Recourse to other assays may be indicated, in order to assess fully
the in vitro activity of certain test chemicals. Acquisition of
test data from a minimum of these 2 standard assays, if optimally
performed, will enable a greater degree of comparability of data
between both independent laboratories and different countries, and
this can only help the efficient detection of possible new human
carcinogens.
In many testing schemes, resources are spread across a large
number of in vitro assays, and this may lead to assays on a new
chemical being conducted suboptimally. The alternative suggested
here is that with 2 well-conducted and common assays, there could
be enhanced comparability and reliability of data, together with an
increase in carcinogen sensitivity. In addition, the Salmonella
assay should be conducted according to what is now regarded as the
best current practice, i.e., employing strains TA1535, 1537, 1538,
98, and 100 in the presence and absence of an induced rat liver S9
mix (10%) (Ames et al., 1975; UKEMS 1983) and also using a
preincubation protocol with strains TA98 and 100 in the case of
initial negative responses. Use of a different source of S9 mix
(e.g., hamster) and additional levels of rat liver S9 mix (e.g., 4%
and 30%) should also be considered (Venitt et al., 1984). In the
case of volatile chemicals, an enclosed test system should be
employed.
The particular cell line employed for cytogenetic assays may be
critical but should not necessarily be restricted to the widely-
used Chinese hamster ovary (CHO) line. Good results were obtained
with the Chinese hamster lung (CHL) line in the CSSTT, but it is
not commercially available at present, and while human lymphocytes
present a sensitive and readily-available source of primary euploid
cells, further studies will be required to demonstrate the general
usefulness of this system (Buckton & Evans, 1982). The Chinese
hamster liver system protocol was designed to measure both
aberrations and aneuploidy and, likewise, will require further
independent studies to confirm the promise shown in the CSSTT.
11. CONCLUSIONS
The dilemma that preceded the formalization of final
conclusions was difficult to resolve. Several carcinogens of
uncertain genotoxicity were selected for the CSSTT, in order to
ascertain which among the many available eukaryotic in vitro
assays for carcinogens would be most efficient and best suited for
adoption as a complement to the generally available Salmonella
mutation assay, by laboratories engaged in the routine screening of
chemicals for potential carcinogens. The organizing committee was
aware of the impact that any conclusions derived from this study
might have on legislative guidelines, but was primarily interested
in the optimal use of the resources of laboratories that may be
attempting to identify possible new human carcinogens. In order to
provide a comprehensive study, the number of types of assays
included was not limited. This non-restrictive method of
collecting assays presented a unique problem; by implication, some
investigators had been invited to submit their assay for scrutiny
for a purpose other than they had originally intended. For
example, many oncologists conduct cell-transformation, aneuploidy,
UDS, etc., assays for research purposes, in order to study the
relationship between genetic end-points and the subtle nuances of
the cancer process. Such research studies may have supported the
selective use of a particular assay that was not claimed to be of
general usefulness for the present purposes, and which was not
being recommended as such by the investigators. It is, therefore,
important to emphasize that the failure of an assay to qualify as a
complementary assay does not preclude its use for research into the
intricate processes of cancer initiation/promotion. This caution
is most important to remember when whole classes of assays such as
"cell transformation", "metabolic cooperation", etc., have been
dismissed from consideration in this collaborative study. On the
one hand, is the immediate and worldwide need for efficient and
reliable means for screening new chemicals, leading to their
registration as safe new products (Ashby et al., 1983); on the
other, the need to use selected in vitro assays to probe the
etiology of chemically-induced cancer. The present study is aimed
solely at the former endeavour, but its findings might expedite the
achievement of the latter. Ancillary findings of the CSSTT are
first, that certain classes of in vitro assays appear to be
insensitive to the cancer-related activities of certain chemical
carcinogens. Second, it is suggested that a clear distinction
should be made between the use of eukaryotic in vitro assays to
confirm genotoxic activity, seen previously for a chemical in
bacteria, and their use to complement bacterial-mutation assays,
i.e., to detect genotoxins found inactive in bacteria; the choice
of assay and the interpretation of data are influenced by the need
originally perceived.
1. Significant differences exist among individual investigators
conducting nominally identical assays. A direct result of the
CSSTT is that many recommended protocol changes were identified
for all assays and some have already been instituted by a
number of investigators. Inadequate test protocols clearly
contributed to the fact that some assays appeared to be
insensitive. It is likely that the need, currently perceived,
to conduct a range of genotoxicity assays on a new chemical has
been stimulated, in part, by the inadequacy of some test
protocols.
2. Urgent attention should be given to the definition of criteria
for judging positive and negative test responses in all assays.
Methods by which weak responses can be interpreted with
confidence were identified, including the adoption of
appropriately sensitive test protocols, the acquisition of a
sufficiently large control data base, and the use of
appropriate statistical methods for data assessment.
3. Carcinogens that are inactive or difficult to detect in the
Salmonella assay fall into 2 distinct groups in this study.
The first include genotoxins that are probably only non-
mutagenic to Salmonella because of protocol deficiencies
associated with the overall metabolic capacity of the assay
system. These agents, HMPA, o-toluidine, safrole, and
acrylonitrile, were detected in most of the eukaryotic assays
studied. Thus, a range of assays can act as limited
complements to the Salmonella mutation assay. The other group
of carcinogens, benzene, DEHP, DES, and phenobarbital,
possessed a more selective range of genotoxic activities, and
none of the assays was selectively sensitive to them. Of the
assays studied, only the induction of chromosomal aberrations,
transformation, gene mutations in mammalian cells, and
aneuploidy in yeast gave encouraging overall performances for
the 8 carcinogens (Conclusion 9).
4. A surprising finding of the CSSTT was the apparent mutagenicity
observed for the 3 carcinogens, benzene, DES, and phenobarbital
in some of the mammalian cell gene-mutation assays. These
activities were not seen by the majority of investigators in
this assay group and were not generally confirmed in either the
Drosophila or fungal gene-mutation assays. If they prove
repeatable, 3 cell-specific gene mutagens will have been
defined. If they are not repeatable, this will emphasize the
need, already perceived by the assay working group, to improve
the protocol and data assessment processes for this class of
assay. The concomitant monitoring of 2 gene loci in a single-
cell line, and the development of appropriate and common data
assessment procedures represent 2 areas for research with
mammalian cell gene-mutation assays. The precise role of these
assays in routine screening programmes requires clarification;
in particular, whether they are to be used to anticipate
possible mammalian carcinogens or possible mammalian mutagens,
or both. The use of negative results in a mammalian cell gene-
mutation assay to negate positive observations made in the
Salmonella assay may be premature.
5. A range of cell-transformation assays was studied, and the
overall performance of several of them was impressive. The
duplicated studies with the Syrian hamster embryo (SHE) assay
and the mouse C3H10T 1/2 assay gave particularly encouraging
results. Nonetheless, these assays are not recommended for
routine screening purposes, at present, because of the
expertise required to carry them out and because of the
problems that remain with clone selection and judgement of when
chemically-induced transformation has occurred.
6. Both of the non-carcinogens showed low incidences of activity
in vitro, several of which were confirmed and most of which
were weak. The clastogenicity of caprolactam for human
lymphocytes and its somatic cell mutagenicity in Drosophila
were particularly clear-cut. The activities observed for these
agents in certain in vitro assays do not, therefore, correlate
with their rodent carcinogenicity, and thus, they represent
false-positive predictions of carcinogenicity. This underlines
the fact that in vitro assays are of use for predicting, rather
than defining carcinogenic activity.
7. Three of the carcinogens, DES, phenobarbital, and DEHP, were
chosen to evaluate the hypothesis that some chemicals can
induce tumours in rodents without first modifying the integrity
of nuclear DNA. Although these chemicals were generally less
active in the tests than the other carcinogens, they each
displayed a range of genotoxic activities in vitro. DEHP
showed least activity, with levels similar to those observed
for the 2 non-carcinogens. Nevertheless, it induced
significant aneuploidy and cell-transformation effects in an
almost total absence of evidence of a direct interaction with
DNA. The data generated for DES confirmed earlier reports that
it is an aneuploidy-inducing and cell-transforming agent, and
the present study also revealed that it was capable of causing
primary damage to DNA; it cannot, therefore, be regarded as
non-genotoxic. Phenobarbital was perhaps the previously best
documented non-genotoxin of those studied, but it has been
defined herein as a gene mutagen, a clastogen, and an
aneuploidy-inducing agent. It is concluded that the phrase
"non-genotoxic" should only be used when a sufficiently large
genotoxicity data base has been collected.
8. Many of the assays evaluated may have an important research
role. However, if their main function is to act as
confirmatory or complementary assays in conjunction with the
Salmonella test, then substantial redundancy among in vitro
eukaryotic genotoxicity assays is evident.
9. In relation to the stated aim of the CSSTT, it is recommended
that resources be applied to the definition of a generally
acceptable and applicable protocol for the conduct of
chromosomal-aberration assays, preferably in an agreed cell
type. Use of this class of assay in conjunction with an
adequate assessment of the mutagenicity of a chemical for
Salmonella should provide an efficient primary screen for
possible new carcinogens. Certain chemicals that induce
tumours in rodents at elevated dose levels will remain
undetected by these tests, as exemplified by DEHP in the
present study. The "detection", in vitro, of such agents may
be achieved by the incorporation of additional assays, but the
price in terms of a reduction in overall specificity and the
increased burden on technical resources may be high.
Consideration of whether the joint inactivity of a chemical in
an adequate Salmonella assay and an adequate in vitro
cytogenetic assay provides evidence of its inactivity with DNA
and, thus, its probable non-carcinogenicity for mammals, at low
dose levels, is indicated. The adoption of a chromosomal-
aberration assay as a common complementary test would have the
additional advantages of allowing easy comparison of data,
ready access to supplementary cytogenetic assays, and the
provision of test data derived using a genetic end-point,
independent of that of the Salmonella (gene-mutation) assay.
REFERENCES
AMES, B.N., MCCANN, J., & YAMASAKI, E. (1975) Methods for
detecting carcinogens and mutagens with the salmonella/mammalian-
microsome mutagenicity test. Mutat. Res., 31: 347-364.
ASHBY, J. (1983) The distinction between genetic and non-genetic
mechanisms in chemical carcinogenesis. In: Zbinden, G., Cohadon,
F., Detaille, J.Y., & Mazué, G., ed. Current problems in drug
toxicity, London, Paris, John Libby Eurotext.
ASHBY, J. & LEFEVRE, P.A. (1983) Genetic toxicology with
formaldehyde and closely related chemicals. In: Gibson, J.E., ed.
Formaldehyde toxicity, Hemisphere Pub.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr, MATTER,
B.E., & SHELBY, M. (1983) The two IPCS collaborative studies on
short-term tests for genotoxicity and carcinogenicity. Mutat. Res.,
109: 123-126.
ASHBY, J., DE SERRES, F.J., DRAPER, M.H., ISHIDATE M., Jr,
MARGOLIN, B.H., MATTER, B.E., & SHELBY, M., ed. (1985) Evaluation
of short-term tests for carcinogens. Report of the International
Programme on Chemical Safety Collaborative Study on in vitro
assays, Amsterdam, Elsevier Science Publishers (Progress in
Mutation Research, Vol. 5).
ATHANASIOU, K. & KRYRTOPOULOS, S.A. (1981) Induction of SCE by
non-mutagenic carcinogens. NATO Adv. Study Inst. Ser A, 40: 557-562
(Chem. abst., 97 18839 (1982).
BARRETT, J.C., WONG, A., & MCLACHLAN, J.A. (1981)
Diethylstilboestrol induced neoplastic transformation without
measurable gene mutation at two loci. Science, 212: 1402-1404.
BUCKTON, K.E. & EVANS, H.J. (1982) Human peripheral blood
lymphocyte cultures. An in vitro assay for the cytogenetic effects
of environmental mutagens. In: Hsu, T.C., ed. Cytogenetic assays of
environmental mutagens, Osman, New Jersey, Allanheld.
CAIRNS, J. (1981) The origin of human cancers. Nature (Lond.),
289: 353-357.
CLEMMENSEN, J. & HJALGRIM-JENSEN, S. (1977) On the absence of
carcinogenicity to man of phenobarbital. In: Statistical studies in
malignant neoplasms, Vol. 5, pp. 38-50.
DEAN, B.J. (1978) Genetic toxicology of benzene, toluene,
xylenes, and phenols. Mutat. Res., 47: 75-97.
DE SERRES, F.J. & ASHBY, J., ed. (1981) Evaluation of short-term
tests for carcinogens: report of the international collaborative
program, North-Holland, New York, Elsevier.
GOLDMACHER, V.S. & THILLY, W.G. (1983) Formaldehyde is mutagenic
for cultured human cells. Mutat. Res., 116: 417-422.
HUFF, J.E. (1982) Caprolactam: condensation of the carcinogenesis
bioassay technical report. Environ. Health Perspect., 45: 199-200.
KLEIN, G. (1981) The role of gene dosage and genetic
transpositions in carcinogenesis. Nature (Lond.), 294: 313-318.
MILVY, P. & WOLFF, M. (1977) Mutagenic studies with
acrylonitrile. Mutat. Res., 48: 217-278.
MOODY, D.E. & REDDY, J.K. (1978) Hepatic peroxisome (microbody)
proliferation in rats fed plasticizers and related compounds.
Toxicol. appl. Pharmacol., 45: 497-504.
PHILLIPS, B.J., JAMES, T.E.B., & GANGOLLI, S.D. (1982)
Genotoxicity studies of di(2-ethylhexyl)phthalate and its
metabolities in CHO cells. Mutat. Res., 102: 297-304.
REDDY, E.P., REYNOLDS, R.K., SANTOS, E., & BARBACID, M. (1982) A
point of mutation is responsible for the acquisition of
transforming properties by the T24 human bladder carcinoma
oncogene. Nature (Lond.), 300: 149-152.
STYLES, J.A (1977) A method for detecting carcinogenic organic
chemicals using mammalian cells in culture. Br. J. Cancer, 36: 558-563.
TABIN, C.J., BRADLEY, S.M., BARGMANN, C.I., WEINBERG, R.A.,
PAPAGEORGE, A.G., SCOLNICK, E.M., DHAR, R., LOWY, D.R., & CHANG,
E.H. (1982) Mechanism of activation of a human oncogene. Nature
(Lond.), 300: 143-149.
TOMITA, I., NAKAMURA, Y., AOKI, N., & INUI, N. (1982)
Mutagenic/carcinogenic potential of DEHP and MEHP. Environ. Health
Perspect., 45: 119-125.
UKEMS (1983) Report of the UKEMS sub-committee on guidelines for
mutagenicity testing, Part I, The United Kingdom Environmental
Mutagen Society.
UNITED STATES PUBLIC LAW (1958) United States Code, title 21,
section 348(c)(3)(A).
US NTP (1980) Benzoin - NTP technical report on the toxicology
and carcinogencity of benzoin, Research Triangle Park, North
Carolina, National Toxicology Program (TR No. 204).
US NTP (1982) Caprolactam - NTP technical report on the
toxicology and carcinogenicity of benzoin, Research Triangle Park,
North Carolina, National Toxicology Program (TR No. 214).
VENITT, S., BUSHELL, C.T., & OSBORNE, M. (1977) Mutagenicity of
acrylonitrile (cyanoethylene) in Escherichia coli. Mutat. Res., 45:
283-288.
VENITT, S., CROFTON-SLEIGH, C., & OSBORNE, M.B. (1984) The hair
dye reagent 2-(2,4-diaminophenoxy) ethanol is mutagenic to
Salmonella typhimurium. Mutat. Res., 135: 31-47.
WEINSTEIN, I.B., WIGLER, M., & STADLER, U. (1976) Analysis of the
mechanism of chemical carcinogenesis of epithelial cell cultures,
Lyons, International Agency for Research on Cancer, (IARC
Scientific Publication No. 12, Screening tests in Chemical
Carcinogenesis).
WEISS, R. (1982) The myconcogene in man and birds. Nature
(Lond.), 299: 9-10.
WILLIAMS, G.M. (1981) Liver carcinogenesis: the role for some
chemicals of an epigenetic mechanism of liver-tumour promotion
involving modification of the cell membrane. Food Cosmet. Toxicol.,
19: 577-583.
